Proceedings of the 31th Meeting
Working Group on Prolamin Analysis
and Toxicity (PWG)
Edited by Peter Koehler
German Research Centre for Food Chemistry, Freising
Verlag Deutsche Forschungsanstalt für Lebensmittelchemie - 2018
Preface
Markus Brandt from Böcker GmbH has been attending the meetings of the Working
Group on Prolamin Analysis and Toxicity (PWG) since many years. Several years ago
he approached me and asked if it would be possible for the Böcker Company to host
the PWG meeting. In 2016, I informed him that this would become reality in 2017 and
Markus and his team started organising the 31st PWG meeting with great enthusiasm.
Together with Karoline Schreiber, Markus Düsterberg and the entire Böcker family, he
succeeded in preparing an unforgettable conference. The meeting was held in the
Lindgart Hotel in Minden, Germany, from 28 to 30 September, 2017. Markus and his
team were present during the entire meeting. As in previous meetings in Germany, the
German Coeliac Society (DZG), namely Judith Glöggler, perfectly managed the
registration of the participants. Apart from the group members the audience comprised
invited speakers, guests from academia, industry, and international coeliac societies.
Representatives from cereal starch producers, producers of gluten-free foods, as well
as manufacturers of kits for gluten analysis and of kits for antibody tests in the
serology of coeliac disease participated from industry. The participants had very
interesting one-and-a-half days of presentations, discussions and networking.
As far as gluten analysis is concerned, the 2017 meeting was focussed in particular on
liquid chromatography/mass-spectrometry (LC-MS) as an evolving technique that is
still waiting for approval in routine analysis up to now. Analytical and clinical work in
the field of coeliac disease (CD), non-coeliac gluten sensitivity (NCGS), wheat allergy
and gluten done in the labs of PWG members as well as results of guests and invited
speakers were presented in 19 talks and lively discussed at the meeting. As usual, legal
and regulatory aspects of gluten analysis and labelling concluded the meeting. A
symposium with presentations looking at the importance of the small intestinal
microbiome for CD and NCGS from different perspectives was organised and
perceived very positively by the audience.
The year 2017 was very challenging for me because I changed my job after serious
problems in and with my former institute, which appears to throw away the long
tradition of cereal research in this instant. This situation made me think of resigning
from the position of chairman of the PWG, but my wife, friends and colleagues
encouraged me to continue. Another consequence was that I am no longer able to
distribute the PWG reference material and it has turned out a real challenge to find an
organisation that is willing to take over this task. I would like to express my special
thanks to Markus Brandt who became a fried during the meeting, to his team, as well
as to Judith Glöggler of DZG for her professional help in the organisation of the
meeting. Finally, I express my gratitude to all friends, colleagues, sponsors and
participants for their inspiration and ongoing support of the PWG and the meeting.
Esslingen, March 2018, Peter Koehler
1. Executive Summary
Nineteen presentations covered all aspects related to gluten, coeliac disease and legal
issues. Starting with food technological aspects and aspects of analytical gluten
tracing, the programme included breeding and genetic aspects of wheat, posttranslational
gluten modification by processing, antibodies and serology, alternative
therapies of coeliac disease, non-coeliac gluten sensitivity, aspects of the pathomechanism
of coeliac disease and the significance of the small intestinal microbiome
in the diet of coeliac disease patients. Finally, legal issues of gluten were discussed.
Analytical session
Seven presentations dealt with analytical challenges of gluten, four of them used liquid
chromatography/mass-spectrometry and this showed the increasing importance of this
technique. Also the question, which antibody would be most suitable to quantitate
gluten in routine analysis was intensely discussed by the participants. It appears to be a
fact that different antibodies yield different results in the quantitation of gluten. More
action is necessary in this field and the PWG was asked to start scientific action
regarding gluten determination by immunochemical methods. Finally, breeding and
genetics were on the agenda. For the first time, an approach using CRISPR/Cas9 to
modify and/or remove gliadins from wheat was presented.
Clinical session
This session included five presentations that started with an overview on gluten in
coeliac disease and non-coeliac gluten sensitivity. A rapid HLA DQ2/DQ8 test was
presented that appears to be a promising tool to perform mass screening of coeliac
disease. A novel therapeutic approach for coeliac disease therapy by specific inhibition
of tissue transglutaminase, a study to get more insights into gluten-specific T cells and
a review of serological studies to diagnose coeliac disease completed the clinical
session.
Symposium: The role of intestinal microbiota in coeliac disease
Three presentations on the role of microorganisms in coeliac disease and non-coeliac
gluten sensitivity were given. In one talk the modification of gluten peptides as
affected by microorganisms was described. The second presentation gave an extensive
overview on the small intestinal microbiome of healthy and coeliac disease patients
and left the question whether the microbiome is the reason for or the consequence of
the disease. Finally, the influence of the food constituents gluten and FODMAPS on
the small intestinal microbiome was discussed. Al in all, the symposium showed that
research on the small intestinal microbiome is an exciting new field in coeliac disease
research.
4. Analytical research reports
Sourdough as leavening agent in gluten-free bread
making
Markus Brandt, Markus Düsterberg
Ernst Böcker GmbH & Co. KG, Minden, Germany
History of sourdough fermentation
Fermentation for processing of cereals is probably in use since at least 12.000 years
[1], and the typical sourdough genera Saccharomyces and Lactobacillus were
identified in fermented cereals excavated in Xinjiang (China) [2]. After being
harvested, cereal grains are in a resting state, hard and dry. Only after comminution or
addition of water and subsequent soaking they can be consumed. Addition of water
activates endogenous enzymes and microbiota – a fermentation that changes the
chemical composition of the grain starts, e.g., proteins and carbohydrates are degraded
[3]. Until World War I, sourdough was the predominant leavening agent for bread
production. Lactobacilli and yeasts are – in variable amounts – the dominating
microorganisms in sourdough and both produce the leavening agent carbon dioxide.
From the 15th century on, top-fermenting brewer’s yeast was introduced for bread
baking and further developed. At the end of 18th century, industrial production of
baker’s yeast on cereal substrates begun. With the introduction of aeration of the mash
in 1870, production of yeast was economically advantageous. Lactic acid bacteria
were used as “protective cultures” for preventing infections of the yeast. With the short
supply of cereals during World War I, molasses as carbon source and malt seedlings as
nitrogen source were introduced as alternatives to cereals [4]. In the 1920ies, malt
seedlings were replaced by ammonia salts because the availability of ammonia
increased strongly after introduction of the Haber-Bosch-process and baker’s yeast
replaced sourdough as main leavening agent for bread production in the following
years. The time, in which bread dough underwent fermentation, decreased from hours
or days to only 30 min proofing time in, e.g., the Chorleywood bread process.
Sourdough fermentation of gluten-free doughs
Sourdough fermentation is one of the strategies proposed to reduce or prevent coeliacimmunogenicity
through food [5], because in contrast to baker’s yeast the
microorganisms in sourdough need to grow in the dough and do not only produce
carbon dioxide from sugar. Growth requires a broad spectrum of available sugars,
amino acids, peptides which are released from the cereal matrix by endogenous or
microbial enzymes. The most traditional technique for sourdough fermentation is
backslopping. This is the inoculation of the raw material with a small amount of dough from a previous successful fermentation. This results in a well-adapted microbiota.
Natural pure cultures (“Reinzucht” starters) are derived from these continuously
propagated fermentations and have stable associations of several lactic acid bacteria
and usually one yeast strain. As an example, Fig. 1 shows the leavening activity of
baker’s yeast and a rice sourdough starter. Volume was measured in a graduated
cylinder. Dough with microbial inactive dried rice sourdough was used as control.
Leavening with sourdough resulted in the same volume as leavening with baker’s
yeast and needed around 1 h more proofing time for bread production.
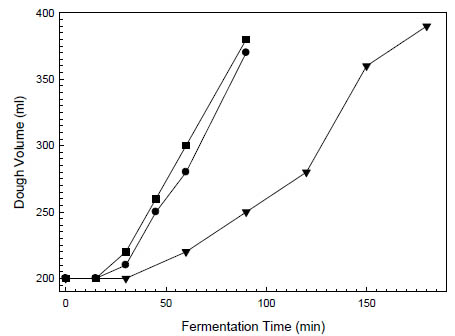
Figure 1. Increase in dough volume of oat bread doughs (dough yield 218) at 28 °C.
The doughs were leavened with (■) 2 % baker’s yeast without dried sourdough, (●)
2 % baker’s yeast with 5 % dried rice sourdough and (▼) 5 % liquid rice sourdough
(Böcker Reinzuchtsauerteig Reis)
The resulting sourdough-leavened oat breads had lower initial hardness (Fig. 2) and a
porosity comparable to the breads leavened with baker’s yeast (data not shown). Other
gluten-free cereals and pseudo-cereals can also be leavened by sourdough, if the
microbiota fits with the carbohydrate composition of the substrate.
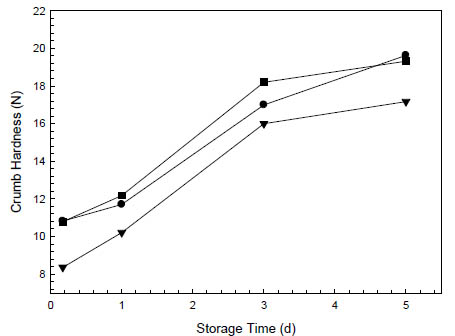
Figure 2. Staling kinetics at 20 °C of oat breads leavened with (■) 2 % baker’s yeast
without dried sourdough, (●) 2 % baker’s yeast with 5 % dried rice sourdough and
(▼) 5 % liquid rice sourdough (Böcker Reinzuchtsauerteig Reis). Hardness was
measured with a SMS TA.XT Plus Texture Analyser with a 12 mm spherical probe at
30 % compression on 2 cm bread slices.
Conclusions
Traditional leavening without baker’s yeast needs longer proofing times than
conventional bread making, on the other hand, a larger amount of the used flour is
subject of the positive effects of fermentation: improvement of flavour, shelf-life or
nutritional composition.
References
1. Hayden B, Canuel N, Shanse J. What was brewing in the Natufian? An
archaeological assessment of brewing technology in the Epipaleolithic. J Archaeol
Method Theory 2013; 20: 102-150.
2. Shevchenkoa A, Yang Y, Knausta A, et al. Proteomics identifies the composition
and manufacturing recipe of the 2500-year old sourdough bread from Subeixi
cemetery in China. J Proteomics 2014; 105: 363-371.
3. Hammes, WP, Brandt MJ, Francis KL, et al. Microbial ecology of cereal
fermentations. Trends Food Sci Technol 2005; 16: 4-11.
4. Kugl J. Der gewerbekundliche Unterricht für Bäcker, Carl Koch, Nuremberg, 1926
5. Gilissen LJWJ, van der Meer IM, Smulders MJM. Reducing the incidence of
allergy and intolerance to cereals. J Cereal Sci 2014; 59: 337-353.
Gluten quantitation by LC-MS: method of choice?
Katharina A. Scherf
Leibniz-Institute for Food Systems Biology at the Technical University of Munich,
Freising, Germany
Introduction
Any quantitative analytical method relies on the measurement of a certain signal, e.g.,
UV absorption, and comparison of the signal to that of a known substance, which
serves as a reference material. This approach works well in case of analytes with a
defined chemical structure, but is not as straightforward in case of polymeric
substances or compound mixtures. Gluten, a highly complex mixture of mono-, oligoand
polymeric proteins, is a challenging analyte, because the composition of gluten
shows great variability depending on species, cultivar, origin and growing conditions
[1]. Despite these issues, analytical methods for gluten should be capable of detecting
all gluten proteins, ideally with comparable specificity and sensitivity. Enzyme-linked
immunosorbent assays (ELISAs) fulfil the requirement of adequate sensitivity with
limits of quantitation down to 3 mg gluten per kg of the food that makes quantitation
of gluten contents possible well below the regulatory threshold of 20 mg/kg for glutenfree
products [2]. Regarding specificity, ELISAs are known to experience some
difficulties due to slight cross-reactivity to oats using the G12 antibody or due to
overestimation of rye and barley in comparison to wheat with the R5 antibody [3].
Furthermore, it is not possible to identify the source of gluten (i.e., wheat, rye or
barley) with the current commercially available ELISA kits. Most antibodies are
specific to certain prolamin types, but do not react to all gluten protein types (Fig. 1) in
a similar way. Despite these disadvantages, ELISAs are fast and comparatively cheap,
suitable for routine analyses and can be performed without specialised equipment.
That is why ELISAs are most commonly used for gluten analysis to assess regulatory
compliance. However, there is no independent protein-based reference method for
verification of ELISA results, although this is an urgent need.
Liquid chromatography-mass spectrometry (LC-MS) is a versatile, specific and
sensitive technique to detect all sorts of analytes including peptides. When using LCMS
for gluten analysis, several points have to be considered. First of all, gluten
proteins have to be extracted and digested to peptides and the choice of enzyme
(trypsin, chymotrypsin or pepsin) has a substantial influence on the results [4]. Then,
the gluten digest is typically cleaned-up prior to analysis using solid phase extraction
or other desalting methods. For LC-MS, there are several choices to be made,
including the instrument itself, data-dependent vs. data-independent acquisition,
untargeted vs. targeted analysis and subsequent data evaluation using bioinformatics
tools [5] and an approach for conversion of peptide contents to gluten content using suitable reference materials. Ideally, LC-MS should detect all coeliac disease (CD)-
immunogenic and toxic peptides known to date, but in case of wheat allergies and noncoeliac
gluten sensitivity other peptides from non-gluten proteins, such as amylasetrypsin-
inhibitors, would have to be considered as well. Several LC-MS methods to
detect CD-active peptides have been published [6-8], but most of those focused on two
to ten marker peptides, a selection that appears rather small compared to the about
1000 known CD-active peptides. Selection criteria for relevant peptides have to be
defined carefully and well-characterised gluten reference materials are essential.
Materials and methods
Isolation of gluten reference materials
Gluten reference materials can be obtained from wheat, rye and barley flours
following the strategy depicted in Fig. 1. After defatting the flours, the albumins and
globulins were removed by extraction with salt solution, the prolamins extracted with
60 % (v/v) aqueous ethanol and the glutelins with 50 % aqueous propanol at pH 7.5
and 60 °C under reducing conditions (1 % dithiothreitol, w/v).
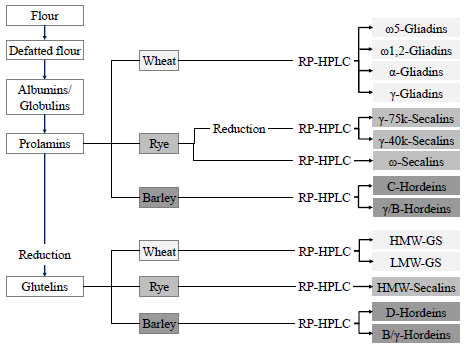
Figure 1. Preparation procedure for gluten protein fractions (prolamins and glutelins)
as well as gluten protein types from wheat, rye and barley flours.
The prolamin and glutelin fractions were concentrated to about half of the original
extract volume using a rotary evaporator, dialysed against distilled water and
lyophilised. The fractions were reconstituted in the respective extraction solvents and
the gluten protein types separated using preparative reversed-phase high performance
liquid chromatography (RP-HPLC). After lyophilisation, the gluten protein types were
profoundly characterised with analytical RP-HPLC, sodium dodecyl sulphate gel
electrophoresis (SDS-PAGE), N-terminal sequencing, LC-MS of the intact proteins
and LC-MS of chymotryptic digests [9]. All methods confirmed that the respective
gluten protein types had been highly enriched. Only B- and γ-hordeins could not be
separated, because they were present in both the prolamin and glutelin fractions and
also had similar retention times on the RP-HPLC column.
Strategy for targeted LC-MS/MS
The purified gluten protein types were used as reference materials for gluten
quantitation by LC-MS/MS. First, the gluten protein types were digested with
chymotrypsin to identify specific peptides that uniquely occur within each protein type
using untargeted LC-MS/MS. Then, the corresponding prolamin and glutelin fractions
and gluten extracts from wheat, rye and barley flours were also analysed to select
peptides that were consistently detected in all three protein isolates (type, fraction and
total gluten extract). From this pool of peptides, the final 16 wheat, 7 rye and 7 barley
marker peptides were required to contain 8-20 amino acids and no cysteine. One 13Cand
15N-labelled peptide standard per cereal species was used that allowed quantitation
by stable isotope dilution assay for the respective peptide and internal calibration for
the other peptides. The most abundant precursor to product ion transitions were
chosen, the collision energies optimised and response factors determined (Fig. 2).
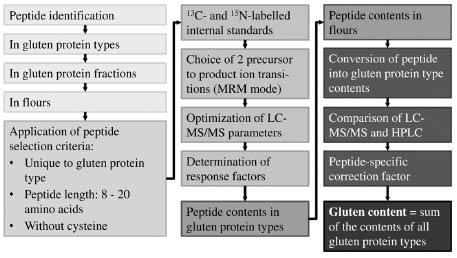

Figure 2. Strategy to develop a comprehensive LC-MS/MS method for gluten from
wheat, rye and barley.
Multiple reaction monitoring (MRM) on a triple-stage quadrupole mass spectrometer
(TSQ Vantage, Thermo Fisher Scientific, Dreieich, Germany) was used in the ESI
positive mode to detect the 30 selected marker peptides and the 3 stable isotope
labelled internal standards. The quantitation of marker peptide contents in gluten
protein types and the gluten extract from flour allowed a conversion of peptide
contents into gluten contents. The results of LC-MS/MS were verified by comparing
the gluten contents to those determined by RP-HPLC and introduction of peptidespecific
correction factors that accounted for losses during digestion. Finally, the
gluten contents of wheat starches and raw materials for rye-based sourdoughs
determined by LC-MS/MS were expressed as sum of all gluten protein types. The LCMS/
MS results were compared to those of other methods, namely R5 ELISA, RPHPLC
and gel permeation HPLC with fluorescence detection (GP-HPLC-FLD) [10].
Results and discussion
The wheat, rye and barley gluten marker peptides included at least one peptide per
gluten protein type to enable comprehensive gluten monitoring also in foods from
which gluten is partially removed during processing (Tab. 1).
Table 1. Amino acid sequences of wheat, rye and barley gluten marker peptides.
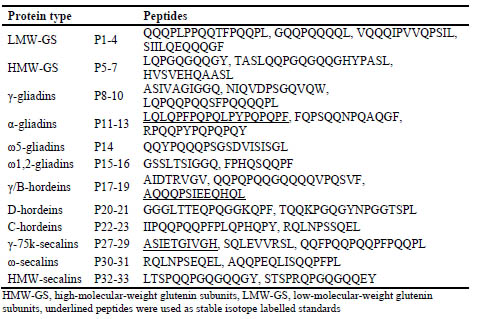
The concentrations of the marker peptides were determined in each gluten protein type
and the results ranged from 0.5 μg/mg ω-secalins (P30) to 48.8 μg/mg D-hordeins
(P21), indicating that some peptide isoforms occurred more frequently than others and
that the efficiency of the enzymatic digest was also highly dependent on the specific
peptide and protein type. The marker peptide concentrations were also analysed in the
gluten extract from the flour mixture and the values lay between 0.1 μg/g flour (P31
from ω-secalins) and 1794 μg/g flour (P17 from γ/B-hordeins), again depending on the
overall content of the protein type within gluten and peptide yields. Peptides P6, P16
and P32 were below the respective limits of detection in the gluten extracts. These
peptide concentrations were used to calculate the content of each protein type in flours.
For verification of the LC-MS/MS results, the content of each protein type determined
by RP-HPLC was defined as 100 % and the recoveries by LC-MS/MS were calculated.
The recoveries ranged from 1.5 % (HMW-secalins determined using P33) to 224 %
(ω1,2-gliadins determined using P15) and also varied for the different peptides that
were derived from the same protein type (e.g., from 64 % to 156 % using either P19 or
P17 to determine γ/B-hordeins). To account for these differences, peptide-specific
correction factors were introduced to ensure 100 % recovery of each protein type.
The LC-MS/MS method together with the final calculation procedure was used to
quantitate gluten in samples of wheat starch (Tab. 2) and raw materials for rye
sourdough fermentation (Tab. 3) as well as the resulting sourdoughs. The gluten
contents determined by LC-MS/MS were expressed as sum of the contents of
individual protein types provided that at least one peptide was detected. In comparison,
the gluten contents determined by RP-HPLC or GP-HPLC-FLD are the sum of
prolamin and glutelin fractions. The gluten contents obtained using ELISA are based
on duplication of the prolamin content as recommended by Codex [2].
Table 2. Contents of wheat marker peptides and resulting contents of gluten protein
types and gluten, respectively, analysed by LC-MS in two wheat starch samples and
comparison of the results to GP-HPLC-FLD and R5 ELISA.
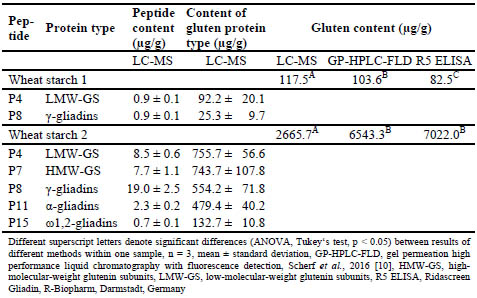
Table 3. Contents of rye marker peptides and resulting gluten contents analysed by
LC-MS in different rye samples and comparison of the results to GP-HPLC-FLD and
R5 ELISA.
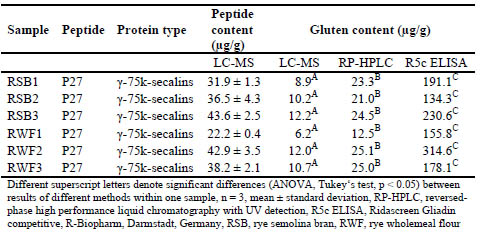
Significant differences between the three analytical methods were found for wheat
starches 1 and 2, rye semolina bran samples 1-3 and rye wholemeal flours 1-3. All
three values for gluten content lay in the same range for wheat starch 1, but the LCMS/
MS values for wheat starch 2 were only about half the values determined by
ELISA and GP-HPLC-FLD. One possible explanation is that other gluten proteins
may have been present that did not contain the specific peptides for which the targeted
LC-MS/MS method is selective. The gluten contents determined in the rye-based raw
materials for sourdough fermentation showed a certain tendency with LC-MS/MS
yielding the lowest and R5 competitive ELISA the highest values. The R5 antibody is
known for its high affinity to rye and resulting overestimation of rye gluten when the
assay is calibrated to wheat [3], so that the high values observed here were not
unexpected. The LC-MS/MS results were about half the values analysed by RP-HPLC,
most likely because only one peptide from γ-75k-secalins was detected and none from
the other rye protein types. All peptides were below the limits of detection in rye
sourdough samples, indicating that they had been hydrolysed during fermentation.
When thinking back to the considerations that have to be made when establishing an
LC-MS method for gluten analysis, the results of this study showed that the conversion
of peptide contents to gluten contents is challenging even if well-defined reference
proteins are available. Furthermore, LC-MS requires expensive and specialised
instrumentation that can only be operated by skilled personnel while method
development and data analysis require a high level of expertise. In case of targeted
LC-MS, only pre-selected peptides are detected and those with amino acid
substitution, deletion or insertion are missed, although they may still be CD-active.
This selectivity for a set of pre-defined peptides is the most likely reason why LC-MS
values tended to be lower in wheat starch and rye samples compared to the other
methods. One major obstacle for LC-MS data evaluation is the lack of curated plant
protein databases that are needed for proper identification of peptide sequences and
linking those back to the intact proteins.
On the positive side, LC-MS is capable of differentiating between wheat, rye, barley
and other cereals, thus allowing the identification of the source of gluten. The method
is applicable to samples from which gluten was partially removed by processing, e.g.,
in beers or wheat starches. Low limits of detection down to 0.03 mg peptide/kg of food
can be achieved using state-of-the-art instrumentation, so that regulatory compliance
may be assessed at the 20 mg/kg level of gluten. It is possible to quantitate specific
peptides very accurately using stable isotope labelled peptide standards. Depending on
the LC-MS instrument, different setups can be used such as various separation
procedures, ionisation techniques and full-scan, single- or multiple-reactionmonitoring
or advanced possibilities like sequential window acquisition of all
theoretical mass spectra. Such untargeted LC-MS analyses generate a huge amount of
data in a comparatively short time that can be evaluated retrospectively in various
ways. As such, LC-MS is a highly versatile technique that can be adapted to different
research questions and offers the possibility of detecting gluten and other allergens in
one multi-analyte run, especially as further advances in LC-MS instrumentation will
contribute to enhancements in selectivity and sensitivity for high-throughput analyses.
Conclusion
The study presented here is the first LC-MS method to use well-defined reference
proteins to detect all gluten protein types from wheat, rye and barley and calculate
peptide-specific yields that can be related back to the intact proteins. The entire
procedure also highlighted the difficulties inherent in this procedure, some of which
can be overcome by using an LC-MS instrument with higher selectivity and sensitivity
and employing stable isotope labelled peptide standards for each marker peptide.
However, some challenges remain like the lack of curated plant protein databases and
the conversion of peptide concentrations to gluten contents, as required by legislation.
Therefore, LC-MS can be seen as a complementary method to ELISA, especially for
samples where ELISA is known to experience difficulties, such as heat-treated or
partially hydrolysed foods.
Acknowledgement The author would like to thank Mrs. Kathrin Schalk and Prof. Dr. Peter Koehler for
their invaluable contributions to the data presented here.
References
1. Hajas L, Scherf KA, Bugyi Z, et al. Variation in protein composition among wheat
(Triticum aestivum L.) cultivars to identify cultivars suitable as reference material
for wheat gluten analysis. Food Chem 2017, doi: 10.1016/j.foodchem.2017.05.005.
2. Codex Standard 118-1979. Codex standard for foods for special dietary use for
persons intolerant to gluten. Codex Alimentarius Commission 2015. Revision 1,
Amendment 2.
3. Lexhaller B, Tompos C, Scherf KA. Fundamental study on reactivities of gluten
protein types from wheat, rye and barley with five sandwich ELISA test kits. Food
Chem 2017, 237: 320-330.
4. Colgrave ML, Byrne K, Howitt CA. Food for thought: Selecting the right enzyme
for the digestion of gluten. Food Chem 2017, 234: 389-397.
5. Colgrave ML, Byrne K, Blundell M, et al. Comparing multiple reaction monitoring
and sequential window acquisition of all theoretical mass spectra for the relative
quantification of barley gluten in selectively bred barley lines. Anal Chem 2016,
88: 9127-9135.
6. Colgrave ML, Goswami H, Byrne K, et al. Proteomic profiling of 16 cereal grais
and the application of targeted proteomics to detect wheat contamination. J
Proteome Res 2015, 14: 2659-2668.
7. Schalk K, Lang C, Wieser H, et al. Quantitation of the immunodominant 33-mer
peptide from α-gliadin in wheat flours by liquid chromatography tandem mass
spectrometry. Sci Rep 2017, 7: 45092.
8. van den Broeck HC, Cordewener JHG, Nessen M, et al. Label free targeted
detection and quantification of celiac disease immunogenic epitopes by mass
spectrometry. J Chrom A 2015, 1391: 60-71.
9. Schalk K, Lexhaller B, Koehler P, Scherf KA. Isolation and characterization of
gluten protein types from wheat, rye, barley and oats for use as reference materials.
PLoS One 2017, 12: e0172819.
10. Scherf KA, Wieser H, Koehler P. Improved quantitation of gluten in wheat starch
for celiac disease patients by gel-permeation high-performance liquid
chromatography with fluorescence detection (GP-HPLC-FLD). J Agric Food Chem
2016, 64: 7622-7631.
Use of LC-MS to detect gluten
Twan America
Plant Breeding, Wageningen University & Research, Wageningen, The Netherlands
No manuscript provided.
Assessing the effect of fermentation and glutenreduction
strategies using LC-MS
Michelle L. Colgrave1, Keren Byrne1, Crispin A. Howitt 2
1 CSIRO Agriculture & Food, 306 Carmody Rd, St Lucia, QLD 4067, Australia
2 CSIRO Agriculture & Food, GPO Box 1700, Black Mountain, ACT 2601, Australia
Introduction
Gluten is the collective name for a class of proteins found in wheat, rye and barley.
Coeliac disease (CD) is an immune-mediated inflammatory disease of the small
intestine in a subset of genetically-susceptible individuals that is triggered by the
ingestion of gluten, resulting in intestinal inflammation and damage. The only current
treatment for CD- and gluten-intolerants (~70 million people globally) is lifelong
avoidance of dietary gluten. Gluten-free (GF) foods are now commonplace, however,
it is difficult to accurately determine the gluten content of GF-products using current
methodologies as the antibodies are non-specific and show cross-reactivity. In
processed products measurement is further confounded by protein modifications
and/or hydrolysis. Gluten measurement in fermented products remains controversial
[1-3]. The industry standard, a competitive ELISA, may indicate gluten values <20
mg/kg, which is deemed safe for people with CD. A recent study on the antibody
response to gluten-reduced beers found that serum from active-CD patients bound to
residual gluten peptides in conventional beers and that a subset of the patient sera
reacted to gluten-removed beers [4]. The authors concluded that residual peptides in
gluten-reduced beers may be specifically recognised by people with CD.
In this study, the hydrolysis of gluten was examined by investigating a range of barleybased
beers crafted to remove gluten using proprietary precipitation and/or application
of commercially-available enzymes, e.g. prolyl endopeptidases (PEP) that degrade the
proline-rich gluten molecules. Proteomic profiling of beers was conducted on the
untreated beers, but also after proteolytic digestion. The beers were also subjected to
size-fractionation to determine the size range of gluten-derived peptide fragments. LCMS
analyses revealed gluten peptides derived from hydrolysed fragments, many >30
kDa in size. Barley gluten (hordeins) were detected in all beers analysed with peptides
representing all hordein classes detected in conventional beers, but also alarmingly in
many gluten-reduced beers. Gluten digestion was incomplete in all commercial beers
employing PEP and peptides comprising missed cleavages were identified [5]
warranting further optimisation of PEP application in an industrial setting.
Materials and methods
Products tested in this study
A selection of beers was purchased internationally from commercial liquor stores
based on their ingredients and gluten status according to their packaging and/or
company website. All beers selected were barley-malt based products rather than
gluten-free beers based on non-gluten containing grains such as rice, sorghum, millet
or tef. A number of regular beers that had previously [6] been shown to contain gluten
were selected as positive controls, C1-C4. The gluten-reduced (or low gluten, LG)
beers, LG1-LG7 and LG9-LG11 are PEP-treated. LG8 is manufactured by an
undisclosed proprietary process. LG12 is brewed with a novel ultra-low gluten barley
[7]. Chemicals, including formic acid (FA), ammonium bicarbonate, dithiothreitol,
iodoacetamide, were purchased from Sigma-Aldrich (Sydney, NSW, Australia).
Acetonitrile were purchased from ChemSupply (Gillman, SA, Australia). Enzymes
used for digestion (trypsin and chymotrypsin) were purchased from Promega (Sydney,
NSW, Australia).
Protein size fractionation, digestion and analysis
Each beer (n=4 replicates per digest) was either: left whole or applied to a 30 kDa or
10 kDa molecular weight cut-off filter as described in [2]. The beers (whole, >30 kDa,
10-30 kDa, <10 kDa) were reduced by addition of DTT and subsequently alkylated by
addition of iodoacetamide (IAM) as described in [5]. To each solution, either trypsin
or chymotrypsin was added and the samples incubated at 37C overnight. The digested
peptide solution was lyophilised and stored at -20C until analysis. The samples were
reconstituted in 100 μL of 1 % formic acid. All beer samples generated were analysed
by LC-MS/MS and proteins identified by automated database searching as described
in [5]. Selected proteins (and their peptide fragments) were further analysed by
targeted proteomics employing multiple reaction monitoring (MRM) MS [5].
Results and discussion
Identification of gluten in control and gluten-reduced beers
Four control beers (C1-C4) that were previously analysed [6] by LC-MS and contained
gluten proteins typical of barley malt beer were analysed. A range of peptides derived
from B-hordeins (examples: I6SJ22; P06470; Q4G3S1), C-hordeins (Q41210;
Q40053), D-hordein (I6TRS8), γ-hordeins (I6TMV6, I6TEV2) and avenin-like
proteins (ALP: F2EGD5, M0VEH1) were detected.
Many of the same proteins (examples: B-hordeins I6SJ22, P06470; C-hordeins
Q40053, Q40037; D-hordein I6TRS8; γ3-hordein I6TEV2; ALP M0VEH1) were
detected in the gluten-reduced beers. Peptides spanning the entire length of the single
gene product D-hordein (I6TRS8) were detected, implying that either the full-length
protein persisted in the gluten-reduced beers or that multiple protein fragments that harboured the detected peptides were present. In order to examine the potential size of
the protein fragments, size fractionation of the beers was undertaken followed by
repeat tryptic digestion and LC-MS/MS analysis. Examining D-hordein (expected MW
75.0 kDa) as detected in a beer crafted to remove gluten (beer LG7), peptides spanning
the entire protein length were detected in both the 10-30 kDa and >30 kDa fractions.
The results were qualitatively similar (LG7, 40.0 % sequence coverage) to that
observed for a control barley beer (C1, 37.9 % sequence coverage) that underwent no
gluten-reduction processes (Fig. 1).
(A) D-hordein as detected in >30 kDa fraction of gluten-reduced beer LG7
MAKRLVLFVAVIVALVALTTAEREINGNNIFLDSRSRQLQCERELQESSLEACRRVVDQQLVGQLPWSTGLQMQC
CQQLRDVSPECRPVALSQVVRQYEQQTEVPSKGGSFYPGGTAPPLQQGGWWGTSVKWYYPDQTSSQQSWQGQQGY
HQSVTSSQQPGQGQQGSYPGSTFPQQPGQGQQPGQRQPWSYPSATFPQQPGQGQGQQGYYPGATSLLQPGQGQQG
PYQSATSPQQPGQGQGQQETYPIATSPHQPGQWQQPGQGQQGYYPSVTSPQQSGQGQQGYPSTTSPQQSGQGQQL
GQGQQPGQGQQGYPSATFPQQPGQWQQGSYPSTTSPQQSGQGQQGYNPSGTSTQQPGQVQQLGQGQQGYYPIATS
PQQPGQGQQLGQGQQPGHGQQLVQGQQQGQGQQGHYPSMTSPHQTGQGQKGYYPSAISPQQSGQGQQGYQPSGAS
SQGSVQGACQHSTSSPQQQAQGCQASSPKQGLGSLYYPSGAYTQQKPGQGYNPGGTSPLHQQGGGFGGGLTTEQP
QGGKQPFHCQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHPGQQTTVSPHQGQQTTVS
PHPGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQPGEQPCGFPGQQTTVSLHH
GQQSNELYYGSPYHVSVEQPSASLKVAKAQQLAAQLPAMCRLEGGGGLLASQ
(B) D-hordein as detected in 10-30 kDa fraction of gluten-reduced beer LG7
MAKRLVLFVAVIVALVALTTAEREINGNNIFLDSRSRQLQCERELQESSLEACRRVVDQQLVGQLPWSTGLQMQC
CQQLRDVSPECRPVALSQVVRQYEQQTEVPSKGGSFYPGGTAPPLQQGGWWGTSVKWYYPDQTSSQQSWQGQQGY
HQSVTSSQQPGQGQQGSYPGSTFPQQPGQGQQPGQRQPWSYPSATFPQQPGQGQGQQGYYPGATSLLQPGQGQQG
PYQSATSPQQPGQGQGQQETYPIATSPHQPGQWQQPGQGQQGYYPSVTSPQQSGQGQQGYPSTTSPQQSGQGQQL
GQGQQPGQGQQGYPSATFPQQPGQWQQGSYPSTTSPQQSGQGQQGYNPSGTSTQQPGQVQQLGQGQQGYYPIATS
PQQPGQGQQLGQGQQPGHGQQLVQGQQQGQGQQGHYPSMTSPHQTGQGQKGYYPSAISPQQSGQGQQGYQPSGAS
SQGSVQGACQHSTSSPQQQAQGCQASSPKQGLGSLYYPSGAYTQQKPGQGYNPGGTSPLHQQGGGFGGGLTTEQP
QGGKQPFHCQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHPGQQTTVSPHQGQQTTVS
PHPGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQPGEQPCGFPGQQTTVSLHH
GQQSNELYYGSPYHVSVEQPSASLKVAKAQQLAAQLPAMCRLEGGGGLLASQ
(C) D-hordein as detected in <10 kDa fraction of gluten-reduced beer LG7
MAKRLVLFVAVIVALVALTTAEREINGNNIFLDSRSRQLQCERELQESSLEACRRVVDQQLVGQLPWSTGLQMQC
CQQLRDVSPECRPVALSQVVRQYEQQTEVPSKGGSFYPGGTAPPLQQGGWWGTSVKWYYPDQTSSQQSWQGQQGY
HQSVTSSQQPGQGQQGSYPGSTFPQQPGQGQQPGQRQPWSYPSATFPQQPGQGQGQQGYYPGATSLLQPGQGQQG
PYQSATSPQQPGQGQGQQETYPIATSPHQPGQWQQPGQGQQGFYPSVTSPQQSGQGQQGYPSTTSPQQSGQGQQL
GQGQQPGQGQQGYPSATFPQQPGQWQQGSYPSTTSPQQSGQGQQGYNPSGTSTQQPGQVQQLGQGQQGYYPIATS
PQQPGQGQQLGQGQQPGHGQQLVQGQQQGQGQQGHYPSMTSPHQTGQGQKGYYPSAISPQQSGQGQQGYQPSGAS
SQGSVQGACQHSTSSPQQQAQGCQASSPKQGLGSLYYPSGAYTQQKPGQGYNPGGTSPLHQQGGGFGGGLTTEQP
QGGKQPFHCQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQTTVS
PHPGQQTTVSPHQGQQTTVSPHQGQQTTVSPHQGQQPGEQPCGFPGQQTTVSLHHGQQSNELYYGSPYHVSVEQP
SASLKVAKAQQLAAQLPAMCRLEGGGGLLASQ
Figure 1. Identification of gluten-derived peptides from D-hordein in glutenreduced
beer, LG7. Protein sequence coverage of D-hordein (I6TRS8) as detected in the >30
kDa size fraction (A); and the 10- 30 kDa fraction (B); revealing peptide
identifications (bold, underlined >95 % confidence) spanning the length of the protein.
Analysis of the <10 kDa fraction (C) that was not subjected to any sample preparation
(reduction, alkylation or digestion) clearly indicated the use of a prolyl endopeptidase
(PEP) in the production of this beer.
The undigested filtrates (<10 kDa) were also analysed revealing a range of internal
gluten peptide fragments. As no digestion was employed, the termini of the peptides
detected may reflect the processes during the brewing of the beers. Many of the
peptides identified in the sub-10 kDa fraction of PEP treated beers were the result of
PEP action with cleavages occurring at P-X, but this was not an efficient process as
there were many “missed” cleavages, that is peptides present that contained X-P-X
motifs (Table 1). The control beers (C1-C4) showed no obvious pattern of protein
cleavage with values ranging from 5-11 % of gluten peptide fragments resulting from
hydrolysis of P-X bonds and >90 % of the gluten peptides detected in C1-C4
contained P-X sites within their sequences. Examining LG1-LG7 and LG9-LG11, the
majority of peptide fragments resulted from cleavage at P-X (52-73 %) indicating the
use of PEP in the production of these beers.
Table 1. Analysis of gluten fragments detected in the sub-10 kDa fraction of control
(C) and low gluten (LG) beers (no enzymatic digestion). The percentage of total
termini that resulted from cleavage at P-X are presented. The number of missed
cleavages (at P-X) within the gluten peptides is also shown.
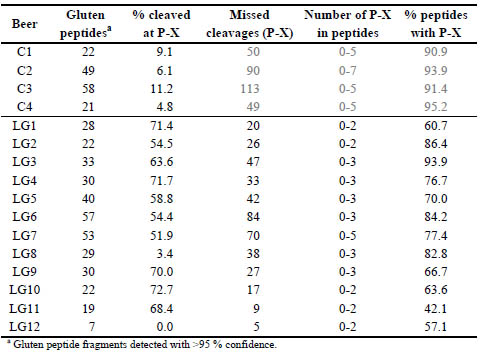
Within the identified gluten peptide sequences in the beers treated by PEP revealed
that 60-94 % contained additional PEP cleavage sites. In fact, only LG11 contained
more completely digested peptides than partially digested (42.1 % missed cleavages).
The untreated beers contained a higher number of P-X sites within an individual
peptide (up to 5 missed cleavages) compared to PEP treated beers where peptides typically only contained 1-2 missed cleavage sites. The exception was LG7 which also
contained a peptide with 5 missed cleavages: QQAELIIP↓QQP↓QQP↓FP↓LQP↓HQP.
Notably, this peptide also contained the QQPFP epitope recognised by the Mendez R5
antibody. For LG8, PEP activity was not apparent with ~3 % of gluten-derived
fragments cleaved at P-X similar to that seen for the control beers. LG8 also yielded a
high proportion of gluten peptide identifications containing P-X sites (82.8 %). As
with all the beers in this study, LG8 was brewed using barley, but the gluten is claimed
to be removed by a proprietary process. Alongside a handful of B- and γ-hordeins, Dhordein
was identified confidently in LG8 by 11 peptides that all clustered in the Cterminal
region of the protein suggesting that a C-terminal fragment persists after
brewing. In LG12, which is a gluten-free beer brewed using a novel gluten-free barley
[7], only γ3-hordein (I6TEV2) was detected by seven peptide fragments resulting from
non-specific cleavage (hydrolysis during brewing).
The beers were sequentially passed through 30 kDa and 10 kDa MWCO filters aimed
at a crude size-based fractionation of the proteins. The retained protein or filtrate for
the sub-10 kDa fraction were then digested with trypsin. Fig. 2 shows the proportion of
the signal as detected in the three size fractions. The B- and γ-hordeins were detected
in both the 10-30 kDa and >30 kDa fractions, which was not a surprising result given
that their expected MW were in the range 28-33 kDa. The D-hordein was most
abundantly detected in the 10-30 kDa fraction (~50 % of D-hordein peptide signal
present, Fig. 6E-H) despite having an expected MW of 75.0 kDa implying that
hydrolysis of the protein had occurred, but this was not dissimilar to that observed for
the control beers where non-specific hydrolysis was also noted to occur.
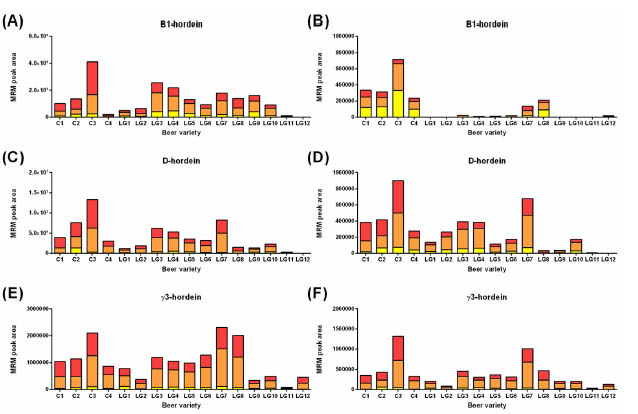
Figure 2. Detection of hydrolysed gluten in size-fractionated beers. The control beers are labelled C1-C4 and the glutenreduced
beers LG1-LG12. The beers were subjected to size-fractionation prior to yield fractions that were expected to contain
proteins >30 kDa (red); 10-30 kDa (orange); and <10 kDa (yellow). Data were analysed by LC-MRM-MS and the integrated
peak area for 6 gluten peptides derived from B1-hordeins (A-B), D-hordein (C-D) and γ3-hordein (E-F) are presented.
Conclusions
The application of PEP during the brewing process reduced the gluten content of beer,
but gluten digestion was not complete in any of the PEP-treated beers analysed with
many missed cleavage sites detected. The treatment with PEP was effective in
reducing the gluten to very low levels in some beers (LG11) and the gluten peptides
that were detected contained a lesser proportion of missed cleavage peptides. The
analysis presented here represents a snap shot in time with only one batch of each beer
tested, but because of the incomplete nature of the treatment it is possible that different
peptides and relative amounts of these peptides would be observed in different
batches.
References
1. Thompson T, Mendez E. Commercial assays to assess gluten content of gluten-free
foods: Why they are not created equal. J Am Diet Assoc 2008; 108: 1682-1687.
2. Colgrave, ML, Goswami H, Blundell M, et al. Using mass spectrometry to detect
hydrolysed gluten in beer that is responsible for false negatives by ELISA. J
Chrom A 2014; 1370: 105-114.
3. Tanner GJ, Colgrave ML, Blundell MJ, et al. Measuring hordein (gluten) in beer– A comparison of ELISA and mass spectrometry. Plos ONE 2013; 8: e56452.
4. Allred LK, Lesko K, Mckiernan D, Kupper C, Guandalini S. The celiac patient
antibody response to conventional and gluten-removed beer. J. AOAC Int. 2017;
100 (2): 485-491.
5. Colgrave ML, Byrne K, Howitt CA. Liquid chromatography-mass spectrometry
analysis reveals gluten in beers crafted to remove gluten. J. Agric. Food Chem.
2017; 65: 9715-9725.
6. Colgrave ML, Goswami H, Howitt CA, Tanner GJ. What is in a beer? Proteomic
characterization and relative quantification of hordein (gluten) in beer. J. Proteome
Res. 2012; 11 (1): 386-396.
7. Tanner GJ, Blundell MJ, Colgrave, ML, Howitt, CA. Creation of the first ultra-low
gluten barley (Hordeum vulgare L.) for coeliac and gluten-intolerant populations.
Plant Biotechnol. J. 2016; 14 (4): 1139-1150.
Influence of food processing on the extractability and
composition of wheat protein fractions
Tanja Schirmer, Katharina Scherf
Leibniz Institute for Food Systems Biology at the Technical University of Munich,
Freising, Germany
Introduction
Currently, immunological methods, i.e. enzyme-linked immunosorbent assays
(ELISA), are recommended by legislation for gluten detection in routine analysis [1].
However, food matrix and the degree of processing can impair gluten quantitation.
Especially, the detection of gluten in heated or extruded products like bread and pasta
[2] and in products containing partially hydrolysed gluten is challenging [3]. Food
processing changes the structure of gluten proteins, which affects their solubility
properties [4] as well as antigen-antibody interactions [5]. The objective of this
research project is to systematically investigate possible causes for the reduced ELISA
sensitivity in processed foods using the bread baking process as model system. The
focus lies on the extractability of different wheat protein fractions from bread (crumb
and crust), analysed by reversed-phase high-performance liquid chromatography (RPHPLC),
as well as on the proteome composition of the fractions, analysed by sodium
dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) and a shotgun
proteomics approach.
Materials and methods
Sample preparation
Kernels of wheat (cv. Akteur, harvested 2015) were milled according to AACCI
Method 26-50.01. The flour was sieved (200 μm) and stored at room temperature (RT)
for 2 weeks. For dough preparation, flour (800 g) was mixed with 3 % baker’s yeast
and 2 % NaCl using a kitchen aid (1 min, level 1). After addition of water (amount
depends on the water absorption which was analysed in a 10 g Farinograph), the dough
was mixed first for 3 min at level 1, then for 3 min at level 3. The dough was placed on
a baking sheet and fermented for 20 min (30 °C, 90 % relative humidity (RH)). About
300 g of dough were hand-shaped, put into a tin pan for proofing for 40 min (at 30 °C,
90 % RH) and baked (25 min at 230 °C). At the beginning of the baking process 2 x
25 mL steam was injected. After baking, the bread was cooled for 2 h at RT, separated
into crumb and crust, lyophilised and milled (6000 rpm, 200 μm sieve) using an Ultra
Centrifugal Mill ZM 200 (Retsch GmbH, Haan, Germany).
Protein extraction
Protein fractions (albumins/globulins, gliadins, glutenins) were extracted using a
modified Osborne fractionation. For analytical purposes (analysis by RP-HPLC),
protein extraction was carried out on a microscale (100 mg; 3 technical replicates)
according to Wieser et al. [6].
For preparative purposes (analysis by SDS-PAGE and LC-MS/MS), protein extraction
was carried out on a macroscale (15 g) according to Schalk et al. [7]. For the latter, the
sample material was defatted with pentane/ethanol (95/5, v/v; 4 x) prior to extraction.
Extracts were lyophilised for further analyses.
Protein content
The protein content (3 technical replicates) of the extracts (from the modified Osborne
fractionation, microscale and macroscale) was determined by RP-HPLC [6].
Lyophilised extracts were re-suspended in their corresponding extraction buffer
(1 mg/mL). The protein content (3 technical replicates) of the sample material and the
extraction sediments was determined according to ICC Standard Method 167 (Dumas
Combustion Principle).
SDS-PAGE
SDS-PAGE was performed according to the protocol described by Lagrain et al. [8].
The amount of lyophilised extracts and extraction sediments analysed corresponded to
1 mg protein/mL extraction buffer and 5 μL were loaded onto the gels.
LC-MS/MS
Lyophilised extracts (1 mg protein/mL; 3 technical replicates) were re-suspended and
chymotryptically hydrolysed according to Rombouts et al. [9]. Protein digests were
purified by solid phase extraction on Discovery® DSC-18 SPE tubes (Sigma-Aldrich
Inc., Steinheim). The eluates were evaporated and re-suspended in 1.0 mL 0.1 %
formic acid for LC-MS/MS analysis. Nanoflow LC-MS/MS was performed by using
an Eksigent nanoLC-Ultra 1D+ system (Eksigent, Dublin, CA) coupled online to a
LTQ-Orbitrap Velos mass spectrometer (Thermo Scientific, Bremen, Germany). 1 μg
of peptides were injected for each measurement. Peptide samples were first loaded on
a trap column (75 μm inner diameter x 2 cm, packed in house with 5 μm, Reprosil
ODS-3; Dr. Maisch, Ammerbuch, Germany) in 100 % loading solvent (0.1 % FA in
HPLC grade water). Peptides were transferred to an analytical column (75 μm x
40 cm, C18 column, Reprosil Gold, 3 μm: Dr. Maisch, Ammerbuch, Germany) and
separated using a 110 min gradient in solvent A (0.1 % FA and 5 % DMSO in HPLC
grade water) and B (0.1 % FA and 5 % DMSO in acetonitrile) from 4-32 % solvent B
at a flow rate of 300 nL/min. MS measurements were performed in data-dependent
acquisition mode, automatically extracting the ten most prominent precursor ions in
the full MS spectra for high energy collision induced dissociation fragmentation at
30 % collision energy. Full MS spectra and MS/MS spectra were acquired at 30.000
resolution and 7.500 resolution, respectively. Dynamic exclusion was set to 60 s.
Label-free quantitation was performed using MaxQuant (version 1.5.3.30) by
searching MS data against a Pooideae database (31.10.2016, 317384 entries) using the
search engine Andromeda. Variable modifications included oxidation of methionine
and N-terminal protein acetylation. Chymotrypsin+ was specified as proteolytic
enzyme with up to two allowed miscleavage sites. Precursor tolerance was set to
10 ppm and fragment ion tolerance was set to 0.05 Da. Label-free quantitation and
match-between-runs options were enabled and results were filtered for a minimal
length of seven amino acids, 1 % peptide and protein false discovery rate as well as
reverse identifications. Statistical analysis (volcano plots) was carried out using
Perseus (version 1.5.6.0).
Results and discussion
The influence of the bread baking process on protein extractability is shown in Fig. 1.
The amount of extracted gliadins was drastically reduced, whereas the amount of
glutenins was increased to a similar extent. The change of the distribution of the
protein fractions is due to the polymerisation of glutenins and gliadins mainly through
heat-induced thiol-/disulphide (SH-SS) interchange reactions [10], triggered by the
baking process. As a result, gliadins were not soluble in aqueous alcohol solutions
anymore and extracted together with the glutenins. Furthermore, the overall
extractability of the total protein was decreased. Thus, the formation of non-SS
crosslinks also needs to be taken into account [11].
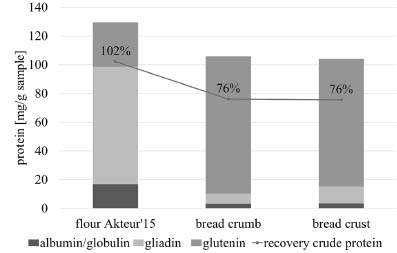
Figure 1. Protein content of different protein extracts as calculated from RP-HPLC
and Dumas data. Differences of all fractions between flour and bread samples are
significant (p < 0.05, ANOVA). Triplicate determination. Standard deviation < 10 %.
The composition of the gliadin and glutenin fractions was investigated in detail by
SDS-PAGE and untargeted LC-MS/MS analysis.
SDS-PAGE was performed to examine the protein band pattern of the lyophilised
extracts and extraction sediments (Fig. 2). In the gliadin fraction, the amount of
proteins containing cysteine residues such as alcohol-soluble high-molecular weight
glutenin subunits (HMW-GS), α- and γ-gliadins was reduced in the crumb and crust
samples in comparison to the flour sample, whereas cysteine-free ω-gliadins
accumulated. These findings confirmed the involvement of cysteine containing
alcohol-soluble proteins in SH-SS interchange reactions during the baking process.
The formation of alcohol-insoluble gluten protein aggregates via SS bonds was
reflected in the glutenin fraction by an increase of certain low-molecular-weight
glutenin subunits (LMW-GS) in the crumb and crust samples. However, the proportion
of HMW-GS as well as of some LMW-GS was decreased in the glutenin fraction
substantiating the assumption that insoluble (non-reducible) gluten protein aggregates
were formed and remained in the extraction sediments of the processed samples. The
SDS-PAGE gel of the sediments demonstrated, that those gluten aggregates could not
be extracted with SDS-buffer either. The extractability of proteins from the crust
sample was remarkably reduced. Thus, the degree of heat treatment seemed to
correlate with the extent of the formation of insoluble gluten protein aggregates.
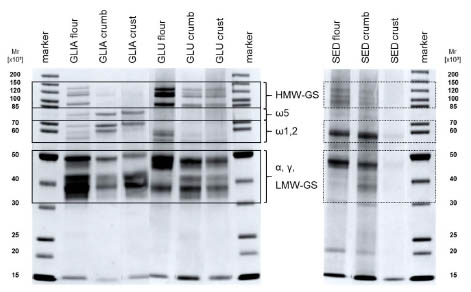
Figure 2. SDS-PAGE of gliadin (GLIA) and glutenin (GLU) fractions isolated from
flour, bread crumb and crust, and their extraction sediments (SED). HMW-GS, LMWGS:
high- and low-molecular-weight glutenin subunits. ω5, ω1,2, α, γ: gliadins.
The proteome of the gliadin and glutenin fractions from flour, crumb and crust
samples was analysed by LC-MS/MS. Tab. 1 comprises all gluten proteins, whose
concentration in the gliadin and/or glutenin fraction from processed samples
significantly (p < 0.05, t-test) changed in comparison to the flour sample.
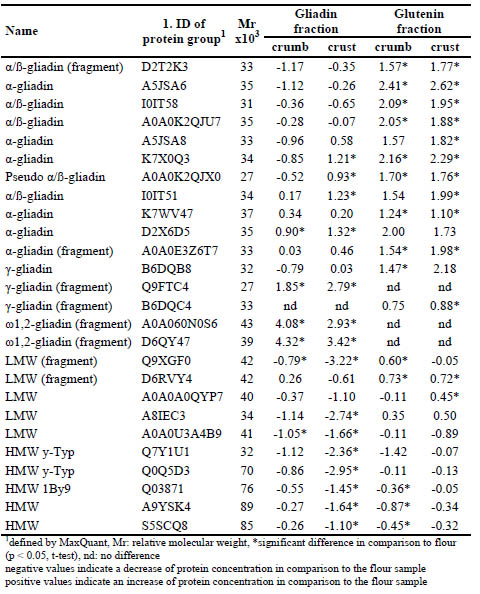
Table 1. Log2-fold change of concentrations of proteins in the gliadin and glutenin
fraction from bread crumb and crust in comparison to flour.
The solubility properties of certain α-gliadins, LMW-GS and HMW-GS were altered.
Seven α-gliadins (D2T2K3, A5JSA6, I0IT58, A0A0K2QJU7, A5JSA8, K7X0Q3,
A0A0K2QJX0) and four LMW-GS (Q9XGF0, D6RVY4, A0A0A0QYP7, A8IEC3)
were present in lower concentrations in the gliadin fraction and in higher
concentrations in the glutenin fraction from crumb and crust samples, i.e. shifted from
the gliadin to the glutenin fractions as seen in the RP-HPLC data. The amount of five
HMW-GS (Q7Y1U1, Q0Q5D3, Q03871, A9YSK4, S5SCQ8) as well as of one LMWGS
(A0A0U3A4B9) was reduced in both fractions from the processed samples. This is
in agreement with the SDS-PAGE results. The LC-MS/MS further revealed, that α-
gliadins and LMW-GS present in the glutenin fractions could not be clearly
distinguished on the SDS-PAGE gel.
The other α-gliadins (I0IT51, K7WV47, D2X6D5, A0A0E3Z6T7), γ-gliadins
(B6DQB8, Q9FTC4, B6DQC4) and especially ω-gliadins (A0A060N0S6, D6QY47)
accumulated in the gliadin and/or glutenin fractions. Beside the lack of cysteine
residues (ω-gliadins), mechanisms why these proteins did not seem to participate in
the gluten network formation need to be further elucidated.
Conclusions
These findings contribute to a better understanding of the influence of food processing
(heat treatment) on the gluten protein structure relevant for ELISA detection. Research
regarding the structure of selected proteins to figure out reaction mechanisms involved
in gluten crosslinking is ongoing.
References
1. Codex Standard 118-1979 Codex Alimentarius Commission 2015
2. Mena MC, Lombardia M, Hernando A, et al. Comprehensive analysis of gluten in
processed foods using a new extraction method and a competitive ELISA based on
the R5 antibody. Talanta 2012; 91: 33-40.
3. Tanner GJ, Colgrave ML, Blundell MJ, et al. Measuring hordein (gluten) in beer -
a comparison of ELISA and mass spectrometry. PLoS One 2013; 8: e56452.
4. Wieser H. Investigations on the extractability of gluten proteins from wheat bread
in comparison with flour. Eur Food Res Technol 1998; 207: 128-132.
5. Mitea C, Kooy-Winkelaar Y, van Veelen P, et al. Fine specificity of monoclonal
antibodies against celiac disease-inducing peptides in the gluteome. Am J Clin Nutr
2008; 88: 1057-1066.
6. Wieser H, Antes S, Seilmeier W. Quantitative determination of gluten protein types
in wheat flour by reversed-phase high-performance liquid chromatography. Cereal
Chem 1998; 75: 644–650.
7. Schalk K, Lexhaller B, Koehler P, et al. Isolation and characterization of gluten
protein types from wheat, rye, barley and oats for use as reference materials. PLoS
One 2017; 12: e0172819.
8. Lagrain B, Rombouts I, Wieser H, et al. A reassessment of the electrophoretic
mobility of high molecular weight glutenin subunits of wheat. J Cereal Sci 2012;
56: 726–732.
9. Rombouts I, Lagrain B, Brunnbauer M, et al. Improved identification of wheat
gluten proteins through alkylation of cysteine residues and peptide-based mass
spectrometry. Sci Rep 3. 2013
10. Schofield JD, Bottomley RC, Timms MF, et al. The effect of heat on wheat gluten
and the involvement of sulphydryl-disulphide interchange reactions. J Cereal Sci
1983; 1: 241–253
11. Rombouts I, Lagrain B, Brijs K, et al. Polymerization reactions of wheat gluten:
the pretzel case. Cereal Foods World 2012; 57: 203–208.
Variations in gluten content estimations revealed by
FAPAS interlaboratory proficiency reports: Dependence
on the use of analytical products, users and range of
concentrations
Miguel Angel Síglez, Carlos Galera, Elena Quesada-Hernández, Ángel Cebolla
Biomedal SL, Seville, Spain
Introduction
Following a gluten-free diet (GFD) is currently the only treatment for coeliac patients.
However, gluten exposure (voluntary or involuntary) is not uncommon, even when
following a GFD. This implies that damage to the gut mucosa as well as very severe
damage to other bodily structures keeps happening even when the patient is
asymptomatic [1,2]. In addition to the obvious reasons why coeliac patients are
exposed to gluten, such as cross-contaminations, omissions, food temptations or
mistakes, the role of the accuracy of analytical techniques to assess the gluten content
in food samples remains unanswered. Food industry players must produce below the
legal limit of 20 mg/kg to be allowed to label their products as “gluten-free” and the
analytical methods they use (mostly immunomethods, ELISA and lateral flow devices)
need to be accurate, repetitive and reproducible.
Currently, there is a wide variety of kits from different manufacturers that differ
mainly in the antibody used. The first generation of polyclonal antibodies (like
the401.21 or Skerrit antibody) which showed limitations in the recognition of certain
gluten types, was then followed by the monoclonal antibody (mAb) R5 [1], which has
a poor recognition of the most immunotoxic peptide, the α-gliadin 33-mer. The last
generation of antibodies comprise the mAbs G12 and A1 [2], which show a strong
recognition of gluten immunogenic peptides (GIP) [3]. Methods based on the R5 mAb
are more widely used than any other, since the sandwich ELISA based on R5 was
endorsed by the Codex Alimentarius as type 1 method [4].
FAPAS is an institution that organizes proficiency tests in which laboratories take part
to verify their quality assurance procedures, monitor their laboratory performances and
compare their methods and results to those of the rest of participants. The data
obtained from every proficiency test are rigorously analysed and a report with
statistical evaluation of results is published by the provider. Those tests of gluten
analysis in different “blind” matrices have been performed since more than a decade.
Laboratories can take part using any of the commercially available ELISA or LF
methods. FAPAS sends out a quality control material with unknown gluten
concentration for the recipient laboratories and subsequently the results submitted are
statistically analysed to provide an assigned value. This assigned value is derived from the consensus of the results obtained by the participant laboratories [5]. Where
possible, FAPAS segregates the results into subsets according to the ELISA kit used.
In this study, we aim to analyse the variability of results obtained by different
analytical kits based on FAPAS reports, and also the results obtained by the most
widely used kit, the Ridascreen Gliadin R5 Sandwich (R-Biopharm), when used by
different laboratories and users.
Materials and methods
We used the reports from FAPAS proficiency tests in which Biomedal’s analytical
laboratory had participated and compared them to the data obtained in our laboratory
by analysing the same FAPAS reference material using the GlutenTox A1/G12
Sandwich ELISA kit (KT-5196, Biomedal, Spain) and the Ingezim Gluten R5
Sandwich ELISA kit (30.GLU.K.2, Ingenasa, Spain). Some data obtained using
Biomedal’s GlutenTox PRO A1/G12 lateral flow kit (KT-5660, Biomedal, Spain) are
also shown.
Results and discussion
Up to 7 FAPAS proficiency tests were analysed (Table 1). Most of the participants
used Ridascreen R5 Sandwich ELISA kit from R-Biopharm (62 participants-75 %),
likely due to the favourable position as Type I method for gluten analysis by Codex
Alimentarius. Despite one of the purposes of a reference method is to contribute to
decrease variations in analytical results, a wide range of values was obtained with this
kit by the different participant laboratories. The minimum values ranged from as low
as 30 % of the assigned value (Ridascreen) to as high as 266 % of the assigned value
(Ridascreen) (Table 1).
Table 1. Analysis of the data obtained after analysing seven FAPAS samples by
different Sandwich ELISA methods
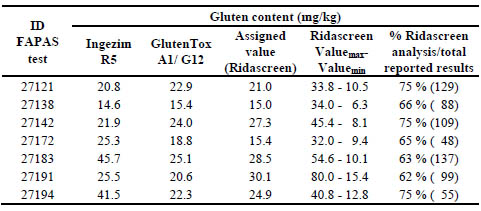
Biomedal analytical laboratory is accredited to the ISO/IEC 17025 standard for its
gluten assay with the Ingezim Gluten R5 Sandwich ELISA kit (Ingenasa) and
participated in those studies using this kit. In parallel, the same reference materials
were analysed by Biomedal using the GlutenTox A1/G12 Sandwich ELISA method
(Biomedal) to study the comparability of the gluten assays. The comparison of the
results for Ingezim Gluten R5 and GlutenTox A1/G12 methods showed less than 25 %
of deviation in 5 out of 7 samples. Furthermore, the results obtained by GlutenTox
A1/G12 kit differed less than 15 % from the assigned value (Ridascreen) in 6 FAPAS
tests, and 32 % in one of them while two results higher than 60 % from the assigned
value were obtained by the Ingezim Gluten R5 kit in two tests.
Another recent FAPAS interlaboratory proficiency test was analysed (report 27204,
Tables 2 and 3) where the GlutenTox PRO A1/G12 lateral flow test kit was assessed in
Biomedal for qualitative estimation of gluten content in two test materials. The
assigned value of each test material calculated from the results obtained by the
Ridascreen R5 Sandwich ELISA test method by FAPAS, TMA24 mg/kg and TMB 41
mg/kg, was correctly estimated by the GlutenTox PRO lateral flow test by using
different cut off by dilutions (>20 mg/kg < 40 mg/kg for TMA; and >40 mg/kg for
TMB). At least one fourth (26.3 %) of the reported values with the Ridascreen kit
might have prompted to the labelling of the analysed material as gluten-free, with less
than 20 mg/kg of gluten (Table 2).
Table 2: Results of collaborative study TMA (Data obtained from FAPAS report
27204 not excluding any result, calculated by Biomedal)
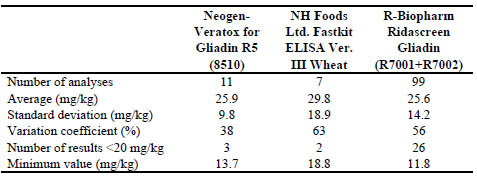
The other laboratory results with other methods (Veratox R5 and NH Food FastKit
ELISAs) had similar proportion of <20 mg/kg reported results (29 % and 27 %,
respectively). The coefficients of variation of the TMA (38 % to 63 %) results were
significantly higher than those of the TMB (37 % to 31 %) when the NH Food FastKit
was used. The lower gluten concentration in a test material, the higher probability of a
variation coefficient increase in the performance of an assay.
Table 3. Results of collaborative study TMB (Data obtained from FAPAS report 27204
not excluding any result, calculated by Biomedal)
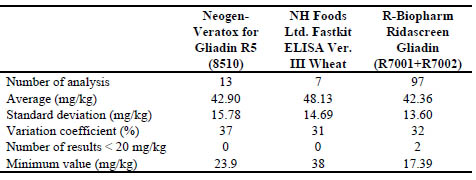
Conclusions
The interlaboratory reproducibility of the analytical results for the same test material
appeared to be more dependent on the different performances by the same user than on
the type of the analytical method used. ELISA methods using different antibodies with
the same laboratory users may produce more similar results than the results obtained
from different laboratories with the same kit. ELISA methods using antibodies other
than R5 antibody can also produce quantitative results of gluten concentration
consistent with the assigned value from the most used ELISA kit.
Coefficient of variation (VC) of the analytical results from the different laboratories
and gluten ELISA kits were considerable higher for gluten concentration close to 20
mg/kg (VC: 38-63 %) than when the gluten concentration was almost two times higher
(VC: <37 %). This high VC in low gluten content close to the legal requirement may
be relevant due to the frequent discrepancies and interpretations of the analytical
results that arise in quality control tests, where the uncertainty of measurement of
every laboratory assay may also play an essential role in food certifications and
labelling. The selection of the same antibody did not solve the problem of
discrepancies in analytical results of same tested material (see also [6]).
Lateral flow immunoassays, as the GlutenTox PRO A1/G12 test kit, could produce
semi quantitative results of gluten concentration in the tested materials consistent with
the assigned value from the most used ELISA kit.
References
1. Torgler C, Siglez MA, Vilchez F, et al. Analytical tools to detect gluten
immunotoxic fractions in food based on monoclonal antibodies raised against the
gliadin 33-mer peptide. In: Stern M (ed): Proceedings of the 24th Meeting of the
Working Group on Prolamin Analysis and Toxicity. Verlag Wissenschaftliche
Scripten, Zwickau, Germany, 2011; pp. 47-50.
2. Morón B, Bethune M, Comino I, et al. Toward the assessment of food toxicity for
celiac patients: Characterization of monoclonal antibodies to a main immunogenic
gluten peptide. PloS One 2008; 3:e2294.
3. Moreno MA, Muñoz-Suano A, López-Casado MÁ, et al. Selective capture of most
celiac immunogenic peptides from hydrolyzed gluten proteins. Food Chem; 2016
205: 36-42.
4. Codex Standard 234-1999, 2014. Recommended methods of analysis and
sampling. Codex Alimentarius commission. Amendment 4.
5. https://fapas.com/
6. Bruins ID, Bremer, MGEG, van der Fels-Klerk I, Hamer RJ. Evaluating the
performance of gluten ELISA test kits: The numbers do not tell the tale. Cereal
Chem; 92: 513-521.
Are we friends with einkorn (Triticum monococcum)?
Gyöngyvér Gell1, Zsófia Birinyi1, Katalin Jäger1, Attila Fábián1, Chris Florides2, Gábor
Veres4, Ilma Rita Korponay-Szabó3, Angéla Juhász1,2
1 Department of Applied Genomics, Agricultural Institute, MTA Centre for Agricultural
Research, Martonvásár, Brunszvik str. 2, H-2462 Hungary
2 Murdoch University, School of Veterinary and Life Sciences Perth WA, Australia
3 Celiac Disease Center, Heim Pál Children's Hospital, Budapest and Department of
Pediatrics, Medical and Health Science Center, University of Debrecen, Debrecen,
Hungary
4 1st Department of Pediatrics, Semmelweis University of Medicine, Budapest,
Hungary
Introduction
Gluten related disorders are a well-studied research area. With the advancement of
cereal diseases, one of the most important goals of the plant research is to identify
setting genotypes of grain with a special flour protein composition beneficial for
patients.
Wheat seed storage proteins are considered as key players in triggering of different
wheat-related health problems. They contain immune responsive peptides (epitopes)
responsible for the autoimmune reaction of coeliac patients.
Today the only effective cure for coeliac disease is a lifelong gluten-free diet.
Although there are several studies published about the disease-triggering proteins in
case of bread wheat, rye or barley, only a limited information is available about the
epitope content, distribution, and frequency of the diploid wheat's, Triticum
monococcum ssp. monococcum and T. monococcum ssp. aegilopoides.
Einkorn (Triticum monococcum ssp. monococcum) is a diploid species of hulled
wheat, with tough glumes ('husks') that tightly enclose the grains. The cultivated form
is similar to the wild form, except that the ear stays intact when ripe and the seeds are
larger [1]. Einkorn is probably the most ancient cultivated wheat species. Its
cultivation started approximately 12000 years ago.
It has much greater genetic variability, than the hexaploid bread wheat. Therefore, it
has more diverse storage protein composition and epitope content as well.
Einkorn wheat has high carotenoid, tocol, micro-element content and decreased
carbohydrate content [2, 3]. It has higher levels of fat, phosphor, sodium, betacarotene,
and pyridoxine than modern wheat species [4]. It can provide easily
digestible food. Therefore, einkorn is a promising candidate for development of bakery
products and has the main role in functional food production in organic farming.
Furthermore due to the simplicity of its genome, einkorn wheat has attracted the interest of the scientific community on nutrition and health aspects in relation to
coeliac disease [5].
The main aim of our research is to characterize the seed proteins of the einkorn
collection of the Cereal Gene Bank of our Institute in Martonvásár using
bioinformatics, proteomics and immunomics analyses. Using a complex workflow, we
can get useful and important information about the toxic protein levels, and
distribution of epitopes of the approximately 200 different einkorn genotypes.
By proteomic and immunological tests, we can identify immune responsive proteins,
which are unsafe even after digestion.
In addition to providing much more accurate information about the relationship
between the storage proteins and the immune response they induce, a large number of
sample suggest, that it is possible to identify some T. monococcum genotypes, that can
be eatable for coeliac patients as well.
Materials and methods
Different Triticum monococcum seeds were derived from the Cereal Gene Bank of the
Department of Plant Genetic Resources and Organic Breeding, Martonvásár.
Patient’s sera suffering from coeliac disease obtained from Dr. Gábor Veres (1st
Department of Pediatrics, Semmelweis University of Medicine) and Dr. Ilma Rita
Korponay-Szabó (Heim Pál Children’s Hospital, Coeliac Centre). In this study, two
coeliac negative, three coeliac disease positive patients on a gluten-free diet and ten
coeliac disease positive sera were used for the estimation of toxic potential the einkorn
seed total protein extract.
In case of T. monococcum total protein extracts, proteins were extracted with SDS
buffer followed the protocol of Dupont and co-workers [6]
Serological ELISA analyses carried out using high binding plates, were coated with
the diluted total protein extracts of the einkorn genotypes (10μg/mL) and incubated
overnight in 4ᵒC degree followed wash steps plates were blocked for 1 h with 5 %
Casein and 0.05 % TWEEN20. The immune reactivity of proteins was detected with
anti-Human IgA (α-chain specific) peroxidase- conjugated antibody produced in goat
(Sigma-Aldrich-A0295) in the presence of 3,3′,5,5′-Tetramethylbenzidine (TMB)
substrate (Sigma-Aldrich, St. Louis, Missouri, United States). For the large-scale,
serological ELISA HLA DQ2.5 and HLA DQ8 human sera of patients with different
age and gender were used in four replicates.
To map the epitopes to the protein sequences coeliac disease-specific linear T-cell and
B-cell epitopes were collected from the ProPepper database [7]. Epitope mapping was
carried out using motif search algorithm of the CLC Genomic Workbench (8.5.1), with
100 % motif identity. In-silico protein digestion was made with PDMQ software tool
[8].
Results and discussion
In our previous study G12 and R5 commercial available ELISA test kits were used and
based on the results of the bioinformatics and immunomics analyses some diploid and
tetraploid species were identified to have significantly lower gliadin content (Fig. 1)
[9]. In one of the investigated T. monococcum genotypes, the toxic protein content
normalised to Chinese Spring bread wheat was significantly lower, so we decided to
make a large scale serological ELISA screen of 200 different wild- and cultivated
einkorn genotypes. There are other studies indicated the lower immunogenicity and
easier digestibility of einkorn bread, but only one or few genotypes were tested [10-
13]. In the first step, the homology test of the seeds was carried out using the gliadin
extracts of the seed endosperms in SDS-PAGE. The seeds of the inhomogeneous genotypes were individually grown in a greenhouse for further analyses.
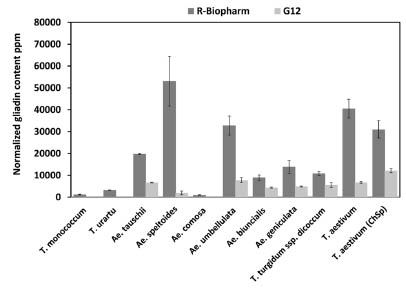
Figure 1. Measured R5 and G12 gliadin contents in diploid, tetraploid and hexaploid
cereal species. Dark grey columns represent the values obtained using the R5 mAb
assay, light grey columns show results for G12 mAb. Error bars represent standard
deviations calculated based on the results of replicates (Gell et al., 2015).
Based on the in-silico epitope mapping T. monococcum sequences retrieved from the
UniProt database all of the alpha-, gamma, -and omega – gliadins containing toxic
epitopes, to some extent resistant to peptic-tryptic digestion (in-silico), but the
composition and the distribution of these epitopes are more diverse than in bread
wheat (Fig 2, Fig 3).
The results of the serological ELISA analyses revealed that from the nearly 200
cultivated and wild einkorn genotypes four have lower toxic protein contents
consequently to Chinese Spring with all of the ten different patient’s sera (Fig 4).
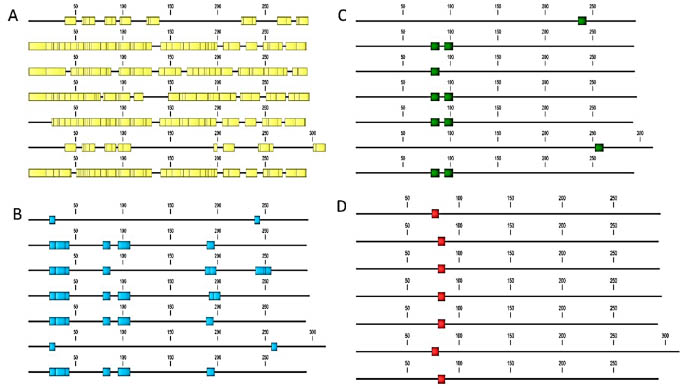
Figure 2. In-silico epitope mapping of T. monococcum α-gliadin sequences retrieved from the UniProt database. A: B cellspecific
epitopes; B: B cell epitopes resistant to proteolytic digestion; C: T cell epitopes; D: T cell epitopes resistant to
proteolytic digestion
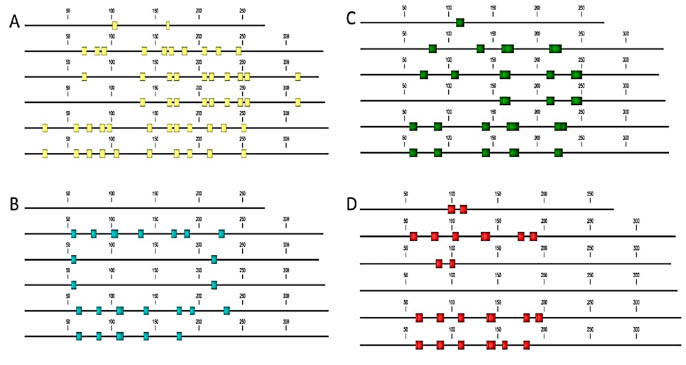
Figure 3. In-silico epitope mapping of T. monococcum ω-gliadins retrieved from the UniProt database. A: B cell-specific
epitopes; B: B cell epitopes resistant to proteolytic digestion; C: T cell epitopes; D: T cell epitopes resistant to proteolytic
digestion
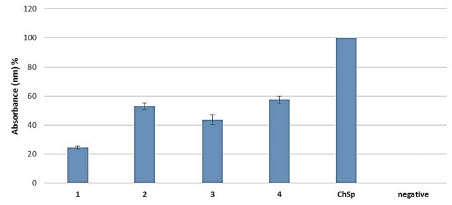
Figure 4. Serological ELISA test of T. monococcum total protein extracts (1 - 4)
compared to Chinese Spring (ChSp) bread wheat with ten different coeliac disease
sera (DQ8 and DQ2).
Compared these results with the commercially available test kits it was revealed that
the R5 and G12 antibody underestimate the toxic protein levels of the T. monococcum.
From one hand the underestimation could be a cause of the variable gliadin
composition and amino acid sequences of einkorn, and from the other hand in the
serological test total extracts were used, could containing non-gluten cross-reactive
proteins.
According to previous examinations connected to climate change with bread wheat,
drought and heat stress and their combinations have a huge impact on the expressed
amounts of storage proteins. Based on these findings further abiotic stress analyses
will be needed to determine the stability of the lower toxic protein content of the
investigated einkorn genotypes.
Conclusions
Due to the large-scale examination of the huge number of einkorn genotypes, highly
diverse population screening has been performed.
Using bioinformatics methods our study confirmed, that einkorn seed storage protein
sequences have more diverse epitope profiles compared to the bread wheat sequences.
In some of the T. monococcum genotypes we have measured significantly reduced
toxic peptide content, however, they were above the officially determined gluten-free
limit. In summary, this result indicates that without further food processing or specific
digestion einkorn genotypes are not suitable food alternatives for coeliac patients.
Further analyses are needed to estimate the environmental effects, abiotic stress factors
on the einkorn seed storage protein composition and toxic epitope expression level.
Therefore it is suggested to perform additional environmental examinations with
einkorn wheat as well.
Funding
This project was supported by OTKA PD 115641 and GINOP-2.3.2-15-2016-00028.
This work was supported by the European Union together with the European Social
Fund (grant number TÁMOP-4.2.4.A/2-11/1-2012-0001 ).
References
1. Cooper R. Re-discovering ancient wheat varieties as functional foods. J Tradit
Complement Med 2015; 5: 138-43.
2. Hidalgo A, Brandolini A, Pompei C, et al. Catotenoids and tocols of einkorn wheat
(Triticum monococcum L.). J Cereal Sci 2006; 44:182-193.
3. Erba D, Hidalgo A, Bresciani J, et al. Environmental and genotypic influences on
trace element and mineral concentrations in whole meal flour of einkorn (Triticum
monococcum L. subsp. monococcum). J Cereal Sci 2011; 54: 250-254.
4. Stallknecht GF, Gilbertson KM, Ranney JM. Alternative wheat cereals as food
grains: Einkorn, emmer, spelt, kamut, and triticale. In: Janick J (ed.), Progress in
new crops. ASHS Press, Alexandria, VA, 1996; pp. 156-170.
5. Agnello PD, Landriscina L, Schiavulli A, et al. Polymeric Proteins Formation
During Pasta-making with Einkorn (Triticum monococcum) and Semolina Mixtures
and Effects on Cooking Behaviour and Acceptability. J Food Proces. Technol
2016; 7: 548-555.
6. Dupont FM, Vensel WH, Tanaka CK, et al. Deciphering the complexities of the
wheat flour proteome using quantitative two-dimensional electrophoresis, three
proteases and tandem mass spectrometry. Proteome Sci 2011; 9: 1-29.
7. Juhász A, Haraszi R, Maulis Cs. ProPepper: a curated database for identification
and analysis of peptide and immune-responsive epitope composition of cereal grain
protein families. Database 2015 bav100, doi:10.1093/database/bav100
8. Haraszi R, Tasi Sc, Juhasz A, et al. PDMQ - Protein Digestion Multi Query
software tool to perform in silico digestion of protein/peptide sequences. BiorXiv,
2015 https://doi.org/10.1101/014019
9. Gell G, Kovács K, Molnár I, et al. Celiac disease specific prolamin peptide content
of wheat relatives and wild species determined by ELISA assays and
bioinformatics analyses. Cereal Res Commun 2015; 43: 133-143.
10. Iacomino G, Di Stasio L, Fierro O, et al. Protective effects of ID331 Triticum
monococcum gliadin on in vitro models of the intestinal epithelium. Food Chem
2016; 212: 537-542.
11. Gianfrani C, Maglio M, Rotondi Aufiero V, et al. Immunogenicity of monococcum
wheat in celiac patients. Am J Clin Nutr 2012; 96: 1339-1345.
12. Pizzuti D, Buda A, D'Odorico A, et al. Lack of intestinal mucosal toxicity of
Triticum monococcum in celiac disease patients. Scand J Gastroenterol 2006; 41:
1305-1311.
13. Zanini B, Villanacci V, De Leo L, et al. Triticum monococcum in patients with
celiac disease: a phase II open study on safety of prolonged daily administration.
Eur J Nutr 2015; 54: 1027-1029.
Gene editing using CRISPR/Cas9 to modify or remove
gliadins from wheat and produce coeliac disease
epitope-free wheat
M. J. M. (René) Smulders1, Aurélie Jouanin1,2, Luud J. W. J. Gilissen3
1 Plant Breeding, Wageningen University & Research, Wageningen, The Netherlands
2 NIAB, Genetics & Breeding, Cambridge, United Kingdom
3 Bioscience, Wageningen University & Research, Wageningen, The Netherlands
Introduction
Cereals cause allergies and intolerances in some people [1]. Individuals that express
the human leukocyte antigen (HLA) DQ2 and/or DQ8 can become intolerant to gluten
proteins from wheat, rye and barley and may develop coeliac disease (CD), a chronic
inflammation of the small intestine. The prevalence of CD is 1-2 % of the general
population worldwide. Strict, gluten-free consumption is currently the only remedy.
Wheat varieties that are safe for people with coeliac disease cannot be produced by
conventional breeding alone [2], as CD epitopes occur in many gliadins and some
glutenin proteins [3], these storage proteins are encoded by gene families with dozens
of members [4,5] partly clustered on two different chromosomes of wheat, and bread
wheat has three sets of chromosomes. Wheat deletion lines, in which parts of
chromosome arms with gluten loci have been eliminated by mutagenesis, are useful
research tools [6], but they also lack many other genes, which may affect their
commercial use [7].
CRISPR/Cas9 is a relatively new technology to perform targeted gene editing, i.e. to
induce mutations and deletions at specific locations in the genome [8-11]. It requires
the introduction into plant cells of a construct that contains the code for a nuclease
(Cas9, or a similar enzyme such as Cpf1) and one or more guide RNA sequences that
target the gene sequence to be altered. The nuclease generates sequence-specific
double strand brakes in the DNA, that will be repaired by the plant but sometimes by
mistake point mutations, indels or other errors are introduced. In wheat this may be
used for the selection of plants with simultaneous mutations in the three genomes [13].
Mutating epitope regions in gluten genes will lower the level of CD epitopes and
ultimately produce CD-safe wheat lines.
In an alternative approach, genetic transformation is avoided. The nuclease protein,
loaded with gRNAs, is added directly to wheat protoplasts or embryogenic cells [16].
This makes the process similar to chemical mutagenesis. However, it is less efficient
for multiple targets than using a transformation step because the added nuclease is
rapidly degraded. It is therefore unlikely that a regenerated plant can be found in
which all extant copies of CD epitopes in gliadin genes have been mutated.
We have edited alpha- and gamma-gliadins of bread wheat using CRISPR/Cas9 with
guide RNAs towards CD epitopes and conserved regions of these gene families, based
on an alignment of many known gliadins. We discuss the results in view of our final
goal of producing a bread wheat variety that has no CD epitopes.
Materials and methods
We stably transformed immature embryos of the spring wheat variety Fielder with
Cas9 + sgRNA constructs directed towards epitopes or conserved regions in the alphagliadins
and gamma-gliadins (Jouanin et al., in prep.). Multiple gRNAs were
combined in a single construct. Transgenic wheat plants (generation T0) were
regenerated in vitro and transferred to a growth chamber for flowering and grain set.
The T1 grains were cut into two parts. One part, containing the embryo, was sown to
produce the T1 plant and further generations. The other part was used for an Acid
PAGE comparison with seeds of the Fielder control and with selected Paragon
gamma-irradiated lines (Jouanin et al., in prep.) and for a proteomics (LC-QTOFMSMS,
[22]) analysis of proteins that were absent and proteins that were mutated in
the gene-edited lines compared to Fielder.
For further evaluation, DNA from the leaves of the corresponding T1 plants will be
extracted to determine the gliadin gene copy number using a quantitative PCR method
(droplet digital PCR), as well as for gluten gene sequencing. As the wheat genome is
very large, an exome capture system was designed that enables isolating and
sequencing only the gluten genes (Jouanin et al., in prep.). The sequence data will give
us additional information about the types of mutations generated by CRISPR/Cas9 in
the gliadins and potentially in off-target sites in the gluten genes.
The wheat lines will be self-pollinated several times to produce plant lines
homozygous for all gliadin mutations and without the CRISPR/Cas9 construct. CD
epitope-specific T cell clones will be used to determine the immunogenicity of the
gluten extracted from the mutant wheat grains. Dough rheology will be measured and,
if necessary, adjusted with, for instance, oat avenins [7] to ensure a decent quality of
bread made using grains from these edited wheat lines. Ultimately, if an edited line
completely devoid of CD epitopes is produced, gluten from the edited wheat line will
be tested on voluntary CD patients in a food challenge, using the prepared bread.
Results and discussion
The Acid PAGE gels show differences in the gliadin protein profiles between the T1
CRISPR grain and the variety Fielder, including disappearance or shifts of bands in
both the alpha-gliadins and the gamma-gliadins, confirming actual mutagenesis.
Interestingly, similar changes are observed in some of the gamma-irradiated lines.
Recently, Sánchez-León and colleagues [13] transformed wheat lines with
CRISPR/Cas9 constructs with gRNAs targeted against alpha-gliadins. They generated plants with mutations in the alpha-gliadin genes, namely small and larger deletions
leading to frameshift mutations and premature stop codons. In gluten extracts from the
T1 grains certain alpha-gliadin bands were absent on Acid PAGE and SDS-PAGE
gels. Surprisingly, in some lines also a decrease in gamma- and omega-gliadins was
observed. Compensatory effects in the production of different gluten gene protein
families was also found in some plants, as previously observed in RNAi lines [14,15].
The total gluten content was up to 85 % lower.
Obviously, both our gene-edited plants and those of Sanchez-León et al. are not yet
safe for CD patients. Not all members of the targeted gliadin families have been
modified, and the epitopes in omega-gliadins and LMW-glutenins [21] were not
targeted at all. In addition, in some of their edited lines Sanchez-León et al. [13]
observed compensation, e.g., higher synthesis of omega-gliadins, which actually
increased the G12 signal. Therefore, we can expect that at least two or three rounds of
CRISPR/Cas9 gene editing will be necessary to modify all epitopes. Alternatively,
parallel modification can be performed for different gluten gene families in different
plants of the same cultivar, after which the modified loci can be combined through
crossing. This may be especially useful when combining safe(r) alpha-gliadin loci on
one or more of the homologous chromosomes 6 with other loci on the homologous
chromosomes 1.
Only a limited number of gliadin genes is expressed at a high level [17]. It is therefore
possible that it may be sufficient to only modify those genes, in all gliadin gene
families to anticipate compensation. However, at this moment we do not know
whether a high level of expression of some genes is completely independent of the
environment, and we also do not know whether the high expression level will
consistently manifest itself in other genotypes after crossing. If so, a mutation that only
modifies an epitope in one of those gliadin proteins, such as a substitution of one or
two amino acids, would be preferable over a mutation that leads to a frameshift
mutation and a truncated mRNA, as this may abolish expression altogether, possibly
leading to compensation by expression of a different gene. For such novel variants we
will be able to predict their immunogenicity, with varying confidence, based on the
knowledge obtained in systematic studies of the immunogenicity of epitope variants
[18] and the 3D structures of the details of the DQ2-epitope-T cell and DQ8-epitope-T
cell protein complexes [19,20]. Tests with T cell clones will need to be done to verify
this.
Whether a gene-edited wheat variety is considered ‘non-transgenic’ in the EU is still
unclear. Since the CRISPR/Cas9-containing construct is introduced by genetic
transformation, some authorities deliberate that the resulting variety may be
considered GM according to the European directive 2001/18/EC, even though the
CRISPR/Cas9 construct is removed by segregation in the offspring of the original
mutated plant (the so-called “null-segregants”) and the variety produced therefore does
not contain any foreign DNA. The absence of foreign DNA also means that the handle
used for safety testing (the transgene, its location, and its possible direct and indirect effects) is absent. Will they forever be considered (according to the process applied) as
transgenic? A product-based safety assessment approach would focus on the traits
introduced in the plant and its safe use, as is the current practice for conventionally
bred plants. Gene-edited varieties are being considered non-GM in other parts of the
world, such as the US, where not the process but the product is evaluated.
Public debates on the use of gene editing technology for improving crop varieties
should include the medical and social benefits associated with using the technology,
i.e., the health benefits of wheat without CD epitopes for the CD patients, the
undiagnosed CD sufferers, and their children. Therefore, CD patients as a prominent
stakeholder group should be involved in the discussion.
A CD-safe wheat line with good baking quality may be introduced in the market as a
specialty variety that has to be produced, processed and sold in a separate production
chain. Such a separate chain will be more difficult to establish, and quality control will
be more challenging, than a regular gluten-free production chain, as the grains and the
plants are morphologically indistinguishable from regular wheat grains and plants. It
would also require an adaptation of the current regulation regarding the 20 ppm gluten
threshold for gluten-free products. Notably, new safety assessment assays would have
to focus on measuring threshold amounts of CD immunogenic epitopes rather than
total gluten.
Conclusions
Mankind has modified many characteristics of wheat during the last 10,000 years,
from non-shattering and hull-less grains in the first millennia to semi-dwarf varieties
with higher yield in the 1960s. Now we have the gene editing technology to also
modify health-impairing plant and food characteristics. Our vision is that of a CD
epitope-free wheat which has largely retained its baking quality. Ultimately, this may
make coeliac disease a very rare disease.
Acknowledgements
The research was partially funded by FP7-PEOPLE-2013-ITN, EID Maximising the
potential of CROP researchers (Max-CROP), grant no. 607178. It does not necessarily
reflect the views of the European Commission and in no way anticipates the
Commission’s future policy in this area.
References
1. Gilissen LJWJ, Van der Meer IM, Smulders MJM. Reducing the incidence of
allergy and intolerance to cereals. J Cereal Sci 2014; 59: 337-353.
2. Jouanin A, Gilissen LJWJ, Boyd LA, et al. Food processing and breeding strategies
for coeliac-safe and healthy wheat products. Food Res Int 2018;
doi.org/10.1016/j.foodres.2017.04.025.
3. Sollid LM, Qiao SW, Anderson RP, et al. Nomenclature and listing of celiac
disease relevant gluten T-cell epitopes restricted by HLA-DQ molecules.
Immunogenetics 2012; 64: 455-60
4. Van Herpen TWJM, Goryunova SV, Van der Schoot J, et al. Alpha-gliadin genes
from the A, B and D genomes of wheat contain different sets of celiac disease
epitopes. BMC Genomics 2006; 7:1.
5. Salentijn EMJ, Mitea DC, Goryunova SV, et al. Celiac disease T-cell epitopes
from gamma-gliadins: immunoreactivity depends on the genome of origin,
transcript frequency, and flanking protein variation. BMC Genomics 2012; 13: 277
6. Van den Broeck HC, van Herpen TWJM, Schuit C, et al. Removing celiac diseaserelated
gluten proteins from bread wheat while retaining technological properties: a
study with Chinese Spring deletion lines. BMC Plant Biol 2009; 9: 41.
7. Van den Broeck HC, Gilissen LJWJ, Smulders MJM, et al. Dough quality of bread
wheat lacking alpha-gliadins with celiac disease epitopes and addition of celiacsafe
avenins to improve dough quality. J Cereal Sci 2011; 53: 206-216.
8. Zhou H, Liu B, Weeks DP, et al. Large chromosomal deletions and inheritable
small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acid Res 2014;
42: 10903-10914.
9. Belhaj K, Chaparro-Garcia A, Kamoun S, et al. Plant genome editing made easy:
targeted mutagenesis in model and crop plants using the CRISPR/Cas system.
Plant Methods 2013; 9: 39.
10. Van de Wiel CCM, JG Schaart, LAP Lotz, MJM Smulders. New traits in crops
produced by genome editing techniques based on deletions. Plant Biotechnol Rep
2017; 11: 1-8.
11. Schaart JG, CCM van de Wiel, LAP Lotz, MJM Smulders. Opportunities for
products of new plant breeding techniques. Trends Plant Sci 2016; 21: 438-449.
12. Wang Y, Cheng X, Shan Q, et al. Simultaneous editing of three homoeoalleles in
hexaploid bread wheat confers heritable resistance to powdery mildew. Nat
Biotechnol 2014; 32: 947-952.
13. Sánchez-León S, Gil-Humanes J, Ozuna CV, Giménez MJ, Sousa C, Voytas DF,
Barro F. Low-gluten, non-transgenic wheat engineered with CRISPR/Cas9. Plant
Biotechnol J 2018; doi: 10.1111/pbi.12837.
14. Gil-Humanes J, Pistón F, Shewry PR, Tosi P, Barro F. Suppression of gliadins
results in altered protein body morphology in wheat. J Exp Bot 2011; 62: 4203-
4213.
15. Becker D, Wieser H, Koehler P, et al. Protein composition and techno-functional
properties of transgenic wheat with reduced alpha-gliadin content obtained by
RNA interference. J Appl Bot Food Qual 2012; 85: 23-33.
16. Liang Z, Chen K, Li T, et al. Efficient DNA-free genome editing of bread wheat
using CRISPR/Cas9 ribonucleoprotein complexes. Nat Commun 2017; 8: 14261.
17. Salentijn EMJ, Esselink DG, Goryunova SV, et al. Quantitative and qualitative
differences in celiac disease epitopes among durum wheat varieties identified
through deep RNA-amplicon sequencing. BMC Genomics 2013; 14: 905.
18. Mitea C, Salentijn EMJ, van Veelen P, et al. A universal approach to eliminate
antigenic properties of alpha-gliadin peptides in celiac disease. PLoS ONE 2010; 5:
e15637.
19. Petersen J, Montserrat V, Mujico JR, et al. T-cell receptor recognition of HLADQ2-
gliadin complexes associated with celiac disease. Nat Struct Mol Biol 2014;
21: 480-488.
20. Petersen J, Kooy-Winkelaar Y, Loh KL, et al. Diverse T cell receptor gene usage
in HLA-DQ8-associated Celiac Disease converges into a consensus binding
solution. Structure 2016; 24: 1643-1657.
21. Sollid LM, Qiao SW, Anderson RP, et al. Nomenclature and listing of celiac
disease relevant gluten T-cell epitopes restricted by HLA-DQ molecules.
Immunogenetics 2012; 64: 455-60.
22. Van den Broeck HC, Cordewener JH, Nessen MA, et al. Label free targeted
detection and quantification of celiac disease immunogenic epitopes by mass
spectrometry. J Chromatogr A 2015; 1391: 60-71.
5. Clinical research reports
Complementarity of 3 monoclonal antibodies (mAbs)
directed against native and deamidated repeat motifs of
gliadins to evaluate the extent of gluten modification
after chemical or enzymatic deamidation processes
Olivier Tranquet1, Valerie Echasserieau-Laporte1, Colette Larré1, Florence Pineau1,
Päivi Kanerva2, Tuula Sontag-Strohm2, Sandra Denery-Papini1
1 UR1268 Biopolymers Interactions Assemblies, INRA, Nantes, France
2 Department of Food and Nutrition, University of Helsinki, Finland
Introduction
Acid-hydrolysed wheat proteins (a-HWP) were used as ingredients in food and
cosmetics. From the 2000’s, cases of severe food allergy to HWP have been reported
in people tolerant to native wheat proteins. More recently in Japan, a soap containing
a-HWP elicited severe skin reactions and food allergy to wheat in over 2000
individuals [1]. Gliadins and glutenin subunits (GS), the main protein fractions of
wheat flour, are characterised by homologous domains consisting of repeated
sequences of 6 to 8 amino acids rich in glutamines. We have shown that deamidation
of these sequences, as the main consequence of acid hydrolysis, generates neoepitopes
responsible for this allergy [2]. The main IgE epitopes involved were
identified for French and Japanese cohorts (QPEEPFPE and PEEPFP respectively)
[2,3]. The detailed analysis of the IgE reactivity of French patients on all the possibilities of deamidation of the peptide QPQQPFPQ revealed that the position and
the extent of deamidation at the scale of this repeated sequence had a strong impact on
the IgE binding. Deamidation of 3 on the 4 glutamines allowed the strongest
IgE/peptide interaction. The determination of the extent of deamidation of wheat
prolamins is essentially based on biochemical methods such as the measurement of
ammonium release after extensive acid hydrolysis [4]. Such a measure gives an overall
result that provides little information about the rate and distribution of deamidation at
the scale of an epitope. A method to assess this rate of deamidation at the epitope
level, together with the presence of residual native repeat peptide would be useful for
the characterisation of glutens modified by chemical or enzymatic processes of
deamidation.
In this work, we evaluated the ability of 3 mouse monoclonal antibodies directed
against native and deamidated repeat motifs of gliadins to discriminate between
gliadin and gluten samples with different deamidation rates.
Material and methods
Antibodies
MAb PQQ3B4 has been raised against the peptide YPQQPFPQ and described in
Tranquet et al. [5]. MAb MCO1 has been raised against the peptide
QPFPQPELPYPQPQ and described in Skovbjerg et al. [6]. Mab INRA-DG1 has been
raised against the peptide LQPEEPFPEQC and described in Tranquet et al. [7]. MAbs
PQQ3B4, and INRA DG1 were used as cell culture supernatant. MC01 was provided
purified by Zedira (Darmstadt, Germany).
Native and deamidated gliadin sub-fractions and total gliadins
Native gluten and native gliadins were previously prepared from wheat flour (c.v.
recital) according to Battais et al. [8]. Briefly, flour was defatted and gluten was
separated from starch by extensive washing of the dough by water. The gliadin
fraction was isolated from the gluten with 70 % ethanol (v/v) and further divided into α, γ, ω2 and ω5-gliadins by ion-exchange chromatography and reverse-phase HPLC.
Three samples of deamidated gliadins with deamidation rates of 15 %, 35 % and 48 %
(referred to as D-GLIA 15, 35 and 48) have been produced by acid hydrolysis of
gliadins and characterised in a previous study [7]. Briefly, Gliadins (10 mg) were
solubilised in 1 mL of 50 % (v/v) ethanol, 0.1N hydrochloric acid, and heated at 90°C
for 40, 90, and 120 min before dialysis against water and lyophilisation. Purified α-, γ, ω2-, and ω5-gliadins, LMW-GS were deamidated by acidic treatment. Level of
deamidation were determined for α-, γ, ω2-, and ω5-gliadins: 52 %, 32 %, 36 % and
51 % respectively [9].
Gluten samples treated by protein glutaminase
Protein-glutaminase (PG) Amano 50 (50 unit/g, 1 % active constituent, 99 % dextrin
as bulking agent), was supplied by Amano Enzyme Inc. (Japan). Vital Gluten (Raisio
Grain Starch Ltd, Finland) at 10 mg/mL was dispersed with PG at 0.13 Unit/mL in
sodium phosphate buffer pH 7 and incubated at 40°C for various periods of time (0-
30h). Control sample (Gluten CTL) was treated under the same conditions without PG
for 24h. After incubation with enzyme, the samples were freeze-dried. In these
conditions, deamidation degrees of 20 %, 45 %, 60 and 72 % were reported for
incubation times of 1 h, 3 h, 5 h, and 30 h respectively [10].
Indirect ELISA
Reactivity of the three antibodies on gliadins and LMW-GS, either native or
deamidated, was assessed by indirect ELISA as described by Battais et al. for sample
preparation and coating [8]. Uncoated sites were blocked with 4 % defatted skimmed
milk in phosphate-buffered saline (PBS) at pH 7.4 for 1 h. MAbs were diluted in PBS
containing 0.1 % skimmed milk (dilution 1:2 for PQQ3B4; MC01 at 0.4 μg/mL;
dilution 1:50 for INRA-DG1) and then added and incubated for 1 h at room
temperature. After washing with PBS 0.05 % Tween 20, bound antibodies were detected with horseradish peroxidase anti-mouse IgG (Bio-Rad, Marnes-la-Coquette,
France, dilution 1:3000) and ortho-phenylenediamine as the substrate. Color
development was stopped with 100 μL of 2 M H2SO4, and the absorbance was read at
492 nm.
Inhibition ELISA
In order to further characterize the reactivity of the 3 mAbs, native or deamidated
gliadin and gluten samples were used as inhibitors in competitive ELISA as described
for INRA-DG1 in Tranquet et al. [7]. For mAb PQQ3B4 (diluted at 1:10), the coating
solution was a total gliadin fraction at 0.3 μg/mL. For the mAb MC01 (diluted at 1 μg/mL), the coating solution was deamidated α-gliadins at 0.5 μg/mL.
Results and discussion
The reactivity of the 3 mAbs on native and deamidated gliadin sub-fractions and
LMW GS were determined by indirect ELISA (Fig 1). Stars indicated the proteins
which contain the immunogenic peptide against which the mAb was raised.
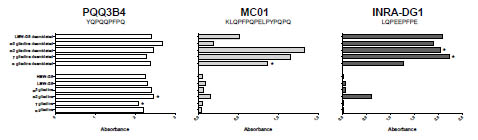
Figure 1. Reactivity of mAbs PQQ3B4, MC01 and INRA-DG1 towards purified native
and deamidated gliadins and GS in indirect ELISA. Stars indicated the proteins in
which the immunogenic peptide was present.
In antigen coated plate ELISA, mAb PQQ3B4 was able to bind to all gliadin classes
and LMW-GS, whether native or deamidated. MAb MC01 recognised deamidated α, γ
and ω2 gliadins and, to a lesser extent, deamidated LMW GS, while deamidated ω5
gliadins, native gliadins and native GS were not bound. INRA-DG1 recognised all
deamidated gliadins and deamidated LMW-GS and showed low but noticeable
reactivity to native ω2 gliadins, while the other native gliadins and GS were not bound.
The three mAbs were able to bind gluten proteins which contain their corresponding
immunogenic peptide (stars in Fig 1) but they also recognised other gliadins and GS.
As gliadins and LMW-GS share homologous sequences in their repetitive domains, the
cross-reactivity of mAbs generated against these sequences was expected and has
already been described [5,11]. Even considering this point, the wide reactivity of mAb
PQQ 3B4 was remarkable. Furthermore, in this assay, the binding of PQQ3B4 to
deamidated gluten proteins raises questions. Was the mAb capable of a certain extentof cross-reactions with deamidated repeated sequences or was its binding only due to
native sequences remaining in acid hydrolysed gluten proteins as previously shown for
mAb R5 [7,12]? Impact of deamidation on mAb PQQ3B4 binding was tested by
pepscan on the peptide QPQQGQQ. The replacement of at least 1 glutamine by
glutamic acid abolished PQQ3B4 binding (data not shown). So it can be assumed that
mAb PQQ3B4 binding on deamidated gliadins was due to interaction with remaining
native epitopes. Both mAbs MC01 and INRA-DG1 have recognised deamidated
gliadins and LMW-GS with minimal cross-reactivity to native proteins. A previous
characterisation had shown that mAb INRA-DG1 has a restricted reactivity and
recognizes strongly deamidated epitopes (XPXEPFPE, where X is Q or E). In the case
of MC01, considering that the peptide against which it was generated has only one
deamidation site, it can be hypothesised that MC01 would rather bind to weakly
deamidated sequences.
In order to go further in determining their ability to discriminate products according to
the presence of native sequences or weakly or strongly deamidated sequences, these
three mAbs, were implemented in competitive ELISA using modified gliadin and
gluten samples whose deamidation rates have been characterised.
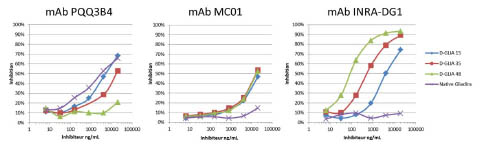
Figure 2a. Characterisation of mAbs PQQ3B4, MC01 and INRA-DG1 in a
competitive ELISA with native and deamidated gliadins (D-GLIA15, 35, 48).
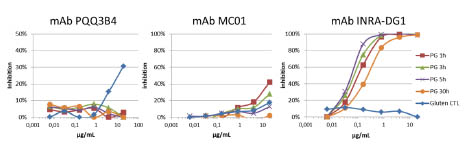
Figure 2b. Characterisation of mAbs PQQ3B4, MC01 and INRA-DG1 in a
competitive ELISA with native and PG-treated gluten 1h, 3h, 5h, 30h.
In the PQQ3B4 competitive ELISA, native gliadins were the most potent inhibitor.
Gliadin samples with low deamidation degree (DD of 15 % and 35 %) were still well
detected by this mAb in contrast to the sample with 48 % of DD. The native gluten
sample also partially inhibited the mAb binding to native gliadins coated on the plate
but none of the PG-treated gluten samples (DD comprised between 20 and 72 %)
inhibited PQQ3B4 mAb binding.
Opposite results were observed in the INRA-DG1 competitive ELISA: native gliadins
or native gluten showed no inhibitory effect, while INRA-DG1 mAb bound well to all
deamidated samples. The inhibitory effect on INRA-DG1 binding gradually increased
as the DD of gliadin and PG-treated gluten samples increased. In competitive ELISA,
the MCO1 mAb bound equally well to the samples D-GLIA 15, 35 and 48, which
displayed an equivalent inhibition capability. In this test, only gluten treated with PG
for 1 or 3 h inhibited MC01 (44 % and 28 % respectively); other PG-glutens were not
detected neither the native samples.
This combination of three competitive ELISAs indicated that during the kinetic of acid
hydrolysis of gliadins, the proportion of native epitopes decreased only significantly
for the sample with a DD of 48 %. Despite different DD (15, 35 and 48 %), similar
amount of weakly deamidated epitopes remained in the three D-GLIA samples; they
much more differ by in their highly deamidated epitopes content. Our combination of
mAbs indicated that only two of the PG-treated samples contained a low amount of
weakly deamidated epitopes while all contained large amount of highly deamidated
epitopes and no native epitopes.
Conclusions
In order to overcome the ambiguity resulting from global determinations of
deamidation rates in deamidated gluten, we evaluated the three mAbs, PQQ3B4,
MC01 and INRA-DG1 in competitive ELISAs for their capacity to differentiate
products based on the extent of deamidation at the epitope level. PQQ3B4 reacts with
native epitopes from the repeated domains of gliadins while MCO1 detect weakly
deamidated epitopes. INRA DG1 binds to highly deamidated sequences which are the
main IgE-binding epitopes involved in allergy to HWP.
References
1. Nakamura M, Yagami A, Hara K, et al. Evaluation of the cross-reactivity of
antigens in Glupearl 19S and other hydrolysed wheat proteins in cosmetics.
Contact Dermatitis 2016; 74: 346–352.
2. Denery-Papini S, Bodinier M, Larré C, et al. Allergy to deamidated gluten in
patients tolerant to wheat: specific epitopes linked to deamidation. Allergy 2012;
67: 1023-1032.
3. Yokooji T, Kurihara S, Murakami T, et al. Characterization of causative allergens
for Wheat-Dependent Exercise-Induced Anaphylaxis sensitized with hydrolyzed
wheat proteins in facial soap. Allergol Int 2013; 62: 435-445.
4. Kanerva PM, Brinck O, Sontag-Strohm TS, et al. Deamidation of gluten proteins
and peptides decreases the antibody affinity in gluten analysis assays. J Cereal Sci
2011; 53: 335-339.
5. Tranquet O, Larré C, Denery-Papini S. Selection of a monoclonal antibody for
detection of gliadins and glutenins: A step towards reliable gluten quantification. J
Cereal Sci 2012; 56:760-763.
6. Skovbjerg H, Koch C, Anthonsen D, et al. Deamidation and cross-linking of
gliadin peptides by transglutaminases and the relation to celiac disease. Biochim
Biophys Acta 2004; 1690: 220-230.
7. Tranquet O, Lupi R, Echasserieau-Laporte V, et al. Characterization of antibodies
and development of an indirect competitive immunoassay for detection of
deamidated gluten. J Agric Food Chem 2015; 63: 5403-5409.
8. Battais F, Pineau F, Popineau Y, et al. Food allergy to wheat: identification of
immunogloglin E and immunoglobulin G-binding proteins with sequential extracts
and purified proteins from wheat flour. Clin Exp Allergy 2003; 33: 962-970.
9. Gourbeyre P, Denery-Papini S, Larré C, et al. Wheat gliadins modified by
deamidation are more efficient than native gliadins in inducing a Th2 response in
Balb/c mice experimentally sensitized to wheat allergens. Mol Nutr Food Res
2012; 56: 336-344.
10. Yong YH, Yamaguchi S, Matsumura Y. Effects of enzymatic deamidation by
protein-glutaminase on structure and functional properties of wheat gluten. J Agric
Food Chem 2006; 54: 6034-6040.
11. Röckendorf N, Meckelein B, Scherf KA, et al. Identification of novel antibodyreactive
detection sites for comprehensive gluten monitoring. PLoS One 2017; 12:
e0181566.
12. Tranquet O, Lupi R, Larré C, et al. Complementarity of native and deamidated
gluten detection with R5 and INRA-DG1 mAbs. In: Koehler P (ed) Proceedings of
the 28th Meeting of the Working Group on Prolamin Analysis and Toxicity.
Deutsche Forschungsanstalt für Lebensmittelchemie, Freising, 2015; pp. 21-25.
Increase in plasma interleukin (IL)-2, IL-8, and IL-10
from 2 to 6 hours after oral gluten challenge observed
in coeliac disease (CD) but not in non-coeliac gluten
sensitivity (NCGS) patients on a gluten-free diet (GFD)
Knut E. A. Lundin1,2 Vikas K. Sarna1,3, Gry I. Skodje1, Suyue Wan3, Leslie J.
Williams4, John L. Dzuri4, Ludvig M. Sollid,1,3 , Robert P. Anderson4
1 KG Jebsen Coeliac Disease Research Centre, University of Oslo, Oslo, Norway
2 Dept of Gastroenterology, Oslo University Hospital-Rikshospitalet, Oslo, Norway
3 Department of Immunology, University of Oslo and Oslo University Hospital-
Rikshospitalet, Oslo, Norway
4 ImmusanT Inc, Cambridge, MA, USA
Introduction
Coeliac disease (CD) is an immune-mediated disease of the small intestine. In patients
with this condition, ingestion of dietary gluten drives an immune response where
CD4+ T cell recognition of deamidated gluten presented by HLA-DQ2 or HLA-DQ8
molecules is of paramount importance [1]. This disease is diagnosed by clinical signs,
serological investigation (IgA against transglutaminase 2) and gastroduodenoscopy
where typical histopathological alterations can be found [2]. Many people in the
society who do not suffer from CD prefer not to eat gluten due to abdominal problems
or feeling of distress. This condition is often referred to as “non-coeliac gluten
sensitivity” or “non-coeliac wheat sensitivity” [2]. It has been claimed that this
condition caused by innate immune response to gluten [3], yet questions have been
raised whether the condition is truly existing [4]. We have done a head-to-head
comparison of gluten challenge in these two clinical entities [5, 6]. Clinical evaluation
of patients with “gluten sensitivity” is important as CD is associated with significant
morbidity and mortality, whereas the NGCS counterpart is not. Importantly, gluten
withdrawal normalizes serology and small intestinal histology in CD, so evaluation of
individuals already on a gluten-free diet is difficult. In common with many active
inflammatory conditions, patients with untreated CD show increased levels of
circulating cytokines that are typically associated with activated lymphocytes and/or
antigen presenting cells [7]. We therefore performed parallel challenges of treated CeD
patients and non-coeliac, but “gluten-sensitive” individuals, who also were on a
gluten-free diet.
Materials and methods
Patients
Two cohorts of patients were studied. CD patients (n=19) had a prior, biopsy-proven
diagnosis in accordance with guidelines [5]. They were recruited from our hospital’s
records, from local support groups or via social media. Prior to challenge, they had a
clinical evaluation, reassessment of gluten-free diet was and a control
gastroduodenoscopy. Only patients with normalised mucosa (mucosal healing of their
diet) could proceed to challenge. NCGS patients (n=49) were also recruited from local
support groups and via social media [6]. None of them had a prior specialist evaluation
of NCGS, but all were on self-instituted gluten-free diet. HLA typing was performed.
Those negative for HLA-DQ2 and HLA-DQ8 were deemed not to have CD. For those
positive for these HLA types, we required a previous gastroduodenoscopy with
negative result while on a gluten containing diet. The studies were approved by the
Regional Ethics Committee and all participants signed an informed, written consent.
Challenge
Both groups were challenged with müsli bars provided by the group of Professor Peter
Gibson, Monash University, Melbourne, Australia. The müsli bars are based on
quionoa, a cereal devoid of gluten and of non-digestible carbohydrates referred to as
FODMAP [8]. The CD patients were challenged, in an open un-blinded manner, with
once daily müsli bar spiked with 5.7 g gluten (without FODMAP) for 14 days [5]. The
NCGS cohort were challenged, in a double-blind, crossover study with un-spiked
müsli bars (placebo), with müsli bar spiked with gluten (same as for the CD patients)
and with müsli bars spiked with 2 g fructans [6]. The results of the challenge outcomes
are reported elsewhere [5, 6].
Symptom scores, immune activation and cytokine measurements
A müsli bar was consumed in the morning with the patient under observation in the
Endoscopy Unit, Oslo University Hospital Rikshospitalet. Symptoms were measured
by GSRS or self-administered visual analog scale. CD subjects were assessed for
serology, duodenal histology, and frequency of gluten-specific T cells in blood using
HLA-DQ:gluten tetramers. The NCGS cohort was followed with symptom scores
only. Venipuncture was done and plasma collected before, and 2, 4 and 6 h after intake
of the müsli bar. Plasma was first analysed for cytokines with a 27-plex cytokine bead
assay from Bio-Rad (Hercules, CA). The advantage of multiplex assays is that they are
labour-efficient and require much less amount of sample, but the kits are expensive.
We used the recommended 1:4 dilution of all samples and found 6 of 27 tested
cytokines to be under the detection limit. The results have been published [5]. We later
analysed plasma cytokines with a more sensitive Mesoscale V-plex assay
(www.mesoscale.com) (manuscript in preparation).
Results
The müsli bars containing gluten induced mucosal changes (villous atrophy as defined
by VH/CrD < 2.0) in 5 of 19 treated CD patients and mobilised HLA-DQ:gluten
tetramer positive T cells in 12 of 15 evaluated patients (Fig. 1).
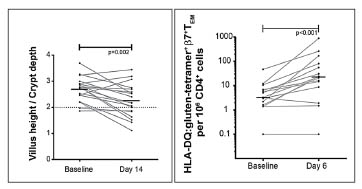
Figure 1. Gluten ingestion for 14 days induced histopathological deterioration as
measured by villous height/crypt depth ratio in the majority of patients (left panel).
HLA-DQ:gluten tetramers could detect immune activation in almost all patients (right
panel). Data published in [5].
In the NCGS patients, there were no signs of symptomatic response to the gluten bars
versus placebo bars (Fig. 2). The symptom scores after FODMAP (fructan) müsli bars
were significantly higher than gluten score for overall GSRS-IBS (p<0.05) and GSRS
bloating (p=0.0003).
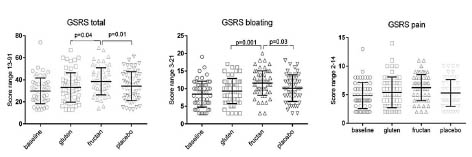
Figure 2. Individuals that reported gluten-sensitivity (but without proper
investigations) were challenged with non-spiked quinoa based müsli bars (placebo) or
spiked with gluten or fructans, respectively. Data published in [6].
Cytokines from challenge of the CD patients were first analysed using the Bio-Rad
multiplex system (Fig. 3).
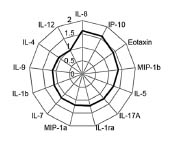
Figure 3. Results of plasma cytokine levels using the Bio-Rad multiplex system. Shown
are fold-change differences compared with baseline levels. Interleukin-2 was also
included in the analysis but the levels were below the assay systems detection limit.
Figure published in [5].
We next analysed the cytokine levels with the more sensitive Mesoscal V-plex assay.
IL-2 fold changes from pre-challenge were significantly increased in CD compared to
NCGS after gluten challenge at 2 h, 4 h and 6 h. Elevations in IL-8 and IL-10 were
also significantly increased in CD compared to NCGS at 4 h and 6 h, but median
elevations were between 1.2 to 1.8-fold.
Discussion
In CD there is response to gluten with concomitant villous blunting, HLA-DQ:gluten
tetramer+ T cells and both “adaptive” and “innate cytokines”. Increase in IL-2, IL-8
and IL-10 after oral gluten challenge is specific for CD, but IL-2 is most sensitive. In
NCGS there is no clinical response to gluten when this is devoid of FODMAP, but
there is clinical response to FODMAP. Measurement of circulating cytokines 4 h after
a single gluten challenge may assist in differentiating between CD and NCGS. It may
be developed as a clinical tool to distinguish the two clinical entities in individuals
already on a gluten-free diet. At present, it should be used in research trials only.
Our findings, and those presented by Tye-Din, Anderson et al., could support a
possible approach to diagnostic workup in patients suspected of having coeliac disease
who have already adopted gluten-free diet. Conceivably such individuals, who have
experienced symptom relief on a gluten-free diet, and who seek investigation of
possible CD, could be a target group for the following work-up: Firstly, HLA typing
should be performed. Those negative for HLA-DQ2 and HLA-DQ8 can, with a high
degree of confidence, be re-assured that they do not have CD. Those who are positive
for HLA-DQ2 or HLA-DQ8, can be offered a single gluten challenge with
measurement of cytokines after 4 h. Whether this challenge needs to be done with “pure” gluten or if it can be done with regular bread, has not been clarified. Those with
a positive cytokine test could be candidates for prolonged gluten challenge followed
by regular clinical investigations [2]. Further refinements to this approach or, alternatively, HLA-DQ: gluten tetramer based diagnostic applications [5,9] may
become parts of the clinician’s toolbox in the future. At this stage, they serve as
research tools only.
References
1. Sollid LM, Iversen R, Steinsbø Ø, et al. Small bowel, celiac disease and adaptive
immunity. Dig Dis 2015; 33: 115-121.
2. Ludvigsson JF, Bai JC, Biagi F, et al. Diagnosis and management of adult coeliac
disease: guidelines from the British Society of Gastroenterology. Gut 2014; 63:
1210-1228.
3. Fasano A, Sapone A, Zevallos V, Schuppan D. Nonceliac gluten sensitivity.
Gastroenterology 2015; 148: 1195-1204.
4. Gibson PR, Skodje GI, Lundin KE. Non-coeliac gluten sensitivity. J Gastroenterol
Hepatol, 2017. 32 (Suppl 1): 86-89.
5. Sarna VK, Lundin KEA, Mørkrid L, et al. HLA-DQ-Gluten tetramer blood test
accurately identifies patients with and without celiac disease in absence of gluten
consumption. Gastroenterology 2017; doi: 10.1053/j.gastro.2017.11.006.
6. Skodje GI, Sarna VK, Minelle IH, et al. Fructan, rather than gluten, induces
symptoms in patients with self-reported non-celiac gluten sensitivity.
Gastroenterology 2017; 154: 529-539.e2.
7. Manavalan JS, Hernandez L, Shah JG, et al. Serum cytokine elevations in celiac
disease: association with disease presentation. Hum Immunol 2010; 71: 50-57.
8. Gibson PR, Shepherd SJ. Food choice as a key management strategy for functional
gastrointestinal symptoms. Am J Gastroenterol 2012; 107: 657-666.
9. Sarna VK, Skodje GI, Reims HM, et al. HLA-DQ:gluten tetramer test in blood
gives better detection of coeliac patients than biopsy after 14-day gluten challenge.
Gut 2017; doi: 10.1136/gutjnl-2017-314461.
Re-exploring the iceberg of coeliac disease in children:
Preliminary results of a multicenter Italian screening
project based on a rapid HLA DQ typing test
Simona Gatti1, Tiziana Galeazzi1, Anil K Verma1, Elisa Franceschini1, Alessandra
Palpacelli1, Giada Del Baldo1, Roberta Annibali1, Antonio Marchesini1, Chiara
Monachesi1, Linda Balanzoni2, Anna M. Colombari2, Maria Teresa Trevisan2, Novella
Scattolo2, Mauro Cinquetti2, Elena Lionetti1, Carlo Catassi1
1 Università Politecnica delle Marche, Ancona, Italy
2 Ospedale "G. Fracastoro", San Bonifacio, Verona, Italy
Introduction
Coeliac disease (CD) is an autoimmune inflammatory process of the small bowel,
related to the ingestion of particular protein mixture “gluten” found mainly in wheat
and similar grains like rye and barley in genetically susceptible individuals [1]. Almost
1 % of the population in Europe and North America are affected with CD [2-4].
Various landmark studies reported a worldwide increase in the prevalence of CD [5-8].
However there are still a high proportion of cases that remain undiagnosed [6]. We
aimed to assess the prevalence of CD autoimmunity and overt CD in Italian school age
children by using HLA typing as the initial screening test, and to redefine the clinical
spectrum of CD.
Materials and methods
Children aged between 5-10 years (49 % males, 51 % females; mean age ±SD: 8.1±0.97
years), attending the primary school from two different regions of Italy, i.e., central
part (Ancona) and Northern part (Verona), were invited to participate. No exclusion
criteria were formulated. Already known CD subjects data were included during the
final study analysis. The diagnostic algorithm of the screening is shown in Fig. 1.
Sample collection
In the school, about 10 μL of blood was collected from each child by a simple prick
test for the HLA DQ determination. Positive HLA DQ2/DQ8 subjects were re-invited
in the respective hospitals, clinically evaluated and 2 mL blood was collected, serum
was separated and kept in -800 C in different aliquots until the different serological
tests were performed.
HLA DQ2/DQ8 typing
HLA DQ2/DQ8 susceptible subjects were identified by the “Celiac Gen Screen Kit” developed by BioDiagene S.R.L (Palermo, Italy), based on a rapid single PCR reactionHLA on a single blood drop. The HLA DQ typing takes about 90 min to complete the
process and includes following steps:
Step 1. Lysis of blood samples collected using EDTA tubes. Ten μL blood sample
was poured in 200 μL of extraction buffer (provided with the Celiac Gene Screen kit)
in 1.5 mL Eppendorf tube and incubated at room temperature. 2 μL of this lysate was
used further for the amplification.
Step 2. Amplification. In 0.2 mL ready to use PCR-tubes provided with the Celiac
Gene Screen kit, 18 μL of ready to use Taq mix (provided with the kit) and 2 μL of
isolated DNA were mixed by pipetting. PCR-tubes were vortex PCR-tubes were
placed into thermo-cycler. After 1.5 h of PCR run, tubes were ready to use for the
analysis.
Step 3. Detection of HLA genes. PCR tubes were placed in the Bio-Run Reader. A
software combines and analyses all data at once and gives the interpretation in ̴ 10– 20 s. Celiac Gene Screen identifies the presence of the alleles that codify for the DQ2
and DQ8 heterodimers that determine the risk of developing CD. depending on the
fluorescence value in the tube, the Bio-Run Reader says that CD associated alleles are
detected or not in the sample.
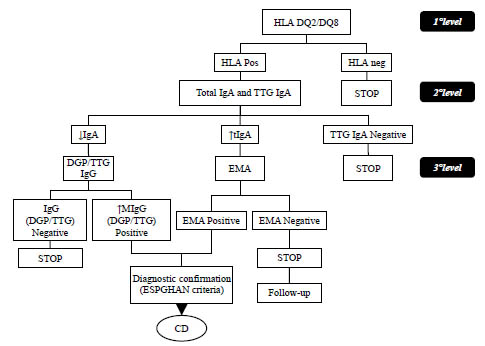
Figure 1. Diagnostic algorithm of the screening. TTG= anti tTG ab, ↑= increased level
of ELISA from the cut off ↓= decreased level of ELISA from the cut off, DGP=
Deamidated gluten peptide, EMA= anti endomysial antibody
Antibody testing
Serum anti-transglutaminase IgA antibodies (TTG) and total IgA were performed in
HLA positive patients. Anti-endomysium antibodies (EMA) and anti-deamidated
gliadin peptides IgG antibodies were searched in TTG positive and IgA deficient
patients respectively. Manufacturer’s guideline was strictly followed for every
serological test. Biopsy was performed according to the ESPGHAN criteria.
Statistical Analysis
Quantitative variables were summarised as mean ± standard deviation (SD),
prevalences and 95 % confidence intervals (CI) were calculated.
Results and discussion
A large number of school children were invited and their screening and further
investigation had been done on the basis of the study algorithm mentioned in the
methodology part. This study has been recently ended; final analysis is still in
progress. At this step we are not able to provide any solid outcome. In coming time
final analysis will be completed and the final result will be revealed.
References
1. Fasano A, Catassi C. Current approaches to diagnosis and treatment of celiac
disease: an evolving spectrum. Gastroenterology 2001; 120: 636-651.
2. Fasano A, Berti I, Gerarduzzi T, et al. Prevalence of celiac disease in at-risk and
not-at-risk groups in the United States: a large multicenter study. Arch Intern Med
2003; 163: 286-292.
3. Maki M, Mustalahti K, Kokkonen J, et al. Prevalence of celiac disease among
children in Finland. N Engl J Med 2003; 348: 2517-2524.
4. Mustalahti K, Catassi C, Reunanen A, et al. Coeliac EU Cluster, Project
Epidemiology. The prevalence of celiac disease in Europe: results of a centralized,
international mass screening project. Ann Med 2010; 42: 587-595.
5. Catassi C, Gatti S, Lionetti E. World perspective and celiac disease epidemiology.
Dig Dis 2015; 33: 141-146.
6. Catassi C, Fasano A. Celiac disease diagnosis: simple rules are better than
complicated algorithms. Am J Med 2010; 123: 691-693.
7. Mustalahti K, Catassi C, Reunanen A, et al. The prevalence of celiac disease in
Europe: results of a centralized, international mass screening project. Ann Med
2010; 42: 587-595.
8. Hill I, Fasano A, Schwartz R, et al. The prevalence of celiac disease in at-risk
groups of children in the United States. J Pediatr 2000; 136: 86-90.
Specific TG2 inhibition and clinical study
Detlef Schuppan
Institut für Translationale Medizin, Johannes-Gutenberg-Universität Mainz, Germany
No manuscript provided.
A clinical trial to dissect the transcriptional programme
of gluten-specific T cells
Stephanie Zühlke1,3, Omri Snir2,3, Shiva Dahal-Koirala2,3, Vikas Kumar Sarna1,4,
Ludvig M. Sollid1,2,3, Knut E. A. Lundin1,4
1 KG Jebsen Coeliac Disease Research Centre, University of Oslo, Norway
2 Center for Immune Regulation, University of Oslo and Oslo University Hospital,
Norway
3 Department of Immunology, University of Oslo, Norway
4 Department of Gastroenterology, Oslo University Hospital, Norway
Introduction
Coeliac disease (CD) is caused by an inappropriate immune response to gluten
proteins found mainly in wheat, rye and barley. Several types of T cells are involved in
CD: a) CD4+ T cells that recognize gluten epitopes presented by HLA-DQ2 or -DQ8
and which orchestrate responses of B cells and other T cells [1, 2], and b)
intraepithelial lymphocytes (IEL) composed of CD8+ αβ T cells and γδ T cells that
induce tissue damage [1, 3].
Concerning gluten specific CD4+ T cells, Anderson et al. have shown that after a
gluten challenge over 3 days in treated CD patients, interferon-γ producing, glutenspecific
cells measured by ELISPOT can be found on day 6 after challenge [4]. Gluten
specific cells CD4+ T cells can also be detected by HLA-DQ:gluten tetramers on day 6
after challenge [5-7]. Notably, HLA-DQ:gluten tetramer positive cells T cells express
gut-homing molecules like β7 integrin and CD103 [5] and the activation marker CD38
[8].
Despite several studies employing HLA-DQ:gluten tetramers [5-7], no kinetic analysis
of appearance of gluten-reactive T cells in blood after gluten challenge has been
performed. In addition, not much is known about functional aspects of HLADQ:
gluten tetramer staining T cells besides their cell surface markers which are
described above.
In this project, we aim to study the kinetics and specific activation of gluten-specific
CD4+ T cells. This can then be used to track down potential therapeutic targets for CD.
Additionally, we will study the frequency of all gut-homing T cells and their response
in blood after gluten challenge.
Materials and methods
CD patients on a gluten-free diet undergo gluten challenge for 3 days with newly inhouse
developed, standardised gluten-containing cookies (see figure 1). Blood samples
are drawn prior to challenge. In the first study (kinetics study), cell kinetics in response to gluten challenge are examined, and additional blood samples are drawn on day 1
and 4 – 8. For the second study (transcriptome study), that will provide insights into
the transcriptomics of gluten-specific T cell, study participants donated blood samples
on day 1 and 6 after the first intake of gluten.
Gluten-specific CD4+ T cells are stained with HLA-DQ:gluten tetramers and for the
kinetics study, their frequency is analysed on flow cytometer for every sample as
described elsewhere [7]. In the transcriptome study, gluten-specific CD4+ T cells
undergo subsequent cell sorting and RNA sequencing [9,10].
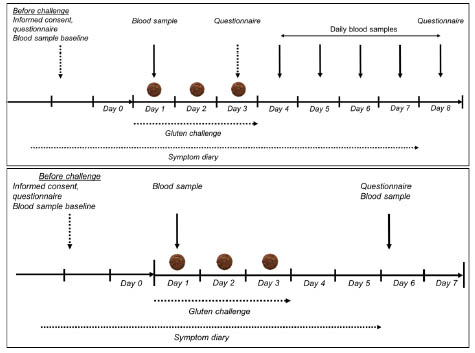
Figure 1. Timeline for the two gluten-challenge studies: the kinetics study (above) and
transcriptome study (below)
The studies are approved by the regional ethical committee of South-East Norway (ref.
2013/1237) and registered at clinicaltrials.gov (NCT NCT02464150).
Results and discussion
In a first round of experiments, we collected blood samples from five participants in
the kinetics study and will in a next step analyse the data. For the transcriptome study,
blood samples from 11 participants have been collected, analysed on flow cytometry
and sorted for RNA-sequencing. All of the patients were challenged with gluten for 3
days.
Data from the first patients indicate a clear efflux of gut-homing gluten-specific T cells
into the blood after challenge.
Conclusions
We are in the ongoing process of patient inclusion for both the kinetics study, as well
as the transcriptome study to achieve a decent number of participants enabling us to
draw reasonable conclusions from our work. The data obtained so far indicate that
gluten challenge with newly developed cookies is provoking a specific T cell response
in blood.
References
1. Sollid LM, Coeliac disease: dissecting a complex inflammatory disorder. Nat Rev
Immunol 2002. 2: 647-55.
2. Sollid LM, Iversen R, Steinsbø Ø, et al. Small bowel, celiac disease and adaptive
immunity. Dig Dis 2015. 33: 115-121.
3. Jabri B, de Serre NP, Cellier C, et al. Selective expansion of intraepithelial
lymphocytes expressing the HLA-E-specific natural killer receptor CD94 in celiac
disease. Gastroenterol 2000. 118: 867-879.
4. Anderson RP, Degano P, Godkin AJ et al. In vivo antigen challenge in celiac
disease identifies a single transglutaminase-modified peptide as the dominant Agliadin
T-cell epitope. Nat Med 2000. 6: 337-342
5. Ráki M, Fallang LE, Brottveit M et al. Tetramer visualization of gut-homing
gluten-specific T cells in the peripheral blood of celiac disease patients. PNAS
2007; 104: 2831-2836.
6. Brottveit M, Ráki, M, Bergseng E et al. Assessing Possible Celiac Disease by an
HLA-DQ2-gliadin Tetramer Test. Am J Gastroenterol 2011. 106: 1318-1324.
7. Sarna VK, Skodje GI, Reims HM, et al. HLA-DQ:gluten tetramer test in blood
gives better detection of coeliac patients than biopsy after 14-day gluten challenge.
Gut 2017; doi: 10.1136/gutjnl-2017-314461.
8. du Pré MF, van Berkel LA, Ráki M et al. CD62LnegCD38+ expression on
circulating CD4+ T cells identifies mucosally differentiated cells in protein fed
mice and in human celiac disease patients and controls. Am J Gastroenterol 2011;
106: 1147-1159
9. Picelli S, Faridani OR, Björklund ÅK et al. Full-length RNA-seq from single cells
using Smart-seq2. Nature protocols 2014. 9: 171-181.
10. http://www.clontech.com/GQ/Products/cDNA_Synthesis_and_Library_Construction
/Next_Gen_Sequencing_Kits/Single_cell_RNA_Seq_Kits_for_mRNA_seq/Single_
Cell_RNA_Seq_v4.
Antibody response to gluten-free diet in paediatric CD
is detectable at three months
David Petroff1, Johannes Wolf2, Thomas Richter3, Marcus K. H. Auth4, Holm H.
Uhlig5,6, Martin W. Laass7, Peter Lauenstein8, Andreas Krahl9, Norman Händel10, Jan
de Laffolie11, Almuthe C. Hauer12, Matthias Heiduk13, Gunter Flemming14, Antje
Schmidt15, Dirk Hasenclever16, Thomas Mothes2
1 Clinical Trial Centre, University of Leipzig, Germany
2 Institute of Laboratory Medicine, Clinical Chemistry and Molecular Diagnostics,
Medical Faculty of the University and University Hospital, Leipzig, Germany
3 Children’s Hospital of the Clinical Centre “Sankt Georg” Leipzig, Germany
4 Alder Hey Children’s National Health Service Foundation Trust, Liverpool, United
Kingdom
5 Translational Gastroenterology Unit, Nuffield Department of Medicine, University of
Oxford, John Radcliffe Hospital, Oxford, United Kingdom
6 Department of Paediatrics, University of Oxford, Oxford, UK
7 University Children’s Hospital, Technical University Dresden, Germany
8 Helios Children’s Hospital, German Clinic for Diagnostics, Wiesbaden, Germany
9 Children’s Hospital “Prinzessin Margaret”, Darmstadt, Germany
10 University Children’s Hospital Leipzig, Germany
11 Children’s Hospital, Justus Liebig University Giessen, Germany
12 University Children’s Hospital Graz, Austria
13 Helios Hospital, Department of Paediatrics, Plauen, Germany
14 Medical School, Hannover, Germany
15 University Children’s Hospital Halle, Germany
16 Institute for Medical Informatics, Statistics & Epidemiology (IMISE), University of
Leipzig, Germany
Introduction
Until recently, histological investigation of mucosal tissue from biopsies was regarded
as a cornerstone of the diagnosis of coeliac disease (CD) [1]. However, it was shown
that assays of IgA-antibodies to tissue transglutaminase (IgA-TTG) in blood samples
are reliable for diagnosis in a large proportion of patients, meaning that the number of
necessary biopsies can be reduced markedly [2-4].
Less is known about the quantitative response of IgA-TTG to a gluten-free diet (GFD).
The concentrations of IgA-TTG generally decrease soon after beginning a GFD, but
rarely normalize within months and remain high in some patients even after years. We
hypothesised that response to GFD can nonetheless be assessed based on short-term
decline in antibody concentrations when measured without upper detection limit.
Materials and methods
This is a secondary, exploratory analysis of data of the prospective antibody study
(AbCD) [3]. The AbCD study enrolled patients aged 5 months to 18 years if a
duodenal biopsy was planned to confirm or refute CD. A baseline blood sample was
tested centrally for IgA-TTG and immunoglobulin G against deamidated gliadin (IgGDGL).
A GFD was recommended if there was strong suspicion of CD. Follow-up
serology was performed blinded about three months later for central antibody
measurements.
To distinguish between a response of antibody concentrations to a GFD from chance
fluctuations, the results of central tests were compared with those of local ones (if
available). The standard deviation of local vs central measurements of both antibodies
in the same patients before GFD was about a factor of two. We define a change of two
standard deviations i.e. a factor of four to be a “substantial response” to GFD.
All central antibody tests were performed with test kits of EUROIMMUN (Lübeck,
Germany). If antibody concentrations were above the measurement range, sera were
serially diluted and values corrected by the dilution factor. For further details see the
paper with the primary results [3].
Results and discussion
From 898 patients, 548 had a GFD and 350 had not. After exclusion of children with
blood samples taken more than 30 days before beginning a GFD, of children initially
on a gluten-reduced, of children without follow-up or with follow-up samples obtained
less than 2 months or later than 6 months after beginning the GFD, of children with
selective IgA-deficiency and of cases with both IgA-TTG and IgG-DGL below the
company cut-off at baseline, 345 children remained for primary analysis (under GFD)
and 46 children for secondary analysis (not under GFD).
The results are summarised in Tab.1. The factor of decrease for IgA-TTG during
short-term GFD is 14. If we consider only the children with a strict diet, the factor is
somewhat higher (15.3). If we consider only children with a less strict diet, the factor
is smaller (5.6). Under normal gluten-containing diet, there are no substantial changes.
In more than 80 % of the children under GFD the concentration of IgA-TTG is
reduced substantially.
The concentration of IgG-DGL decreases during short term-GFD, too. The factor of
decrease is smaller than for IgA-TTG. In 28.1 % of children, the concentration is
reduced more than fourfold.
Of note, even in case of substantial antibody decrease, IgA-TTG remained above the
upper limit of normal in 84 % and above ten times the upper limit of normal
(10×ULN) in 27 %, IgG-DGL remained above the upper limit of normal in 38 % and
above 10×ULN in 5 % of patients.
A limitation of our study is that we have a lack of long-term data and follow-up
biopsies. As a result, we could not follow the progress of “non-responders” nor could
we correlate the antibody response with morphological changes in the intestine. A
second limitation is that we could not perform a gluten challenge, a third limitation,
that it was not possible to assess GFD adherence with an established instrument. We
only could use the data provided by the trial paediatrician who could make a cross
either for strict, less strict or no diet. Otherwise, assessment of GFD compliance was
done according to good clinical practise and the recommendations of the countryspecific
guidelines for management of CD. Finally, our data are only valid for the test
kit of EUROIMMUN.
Table 1. Short-term response of IgA-TTG and of IgG-DGL to gluten-free diet
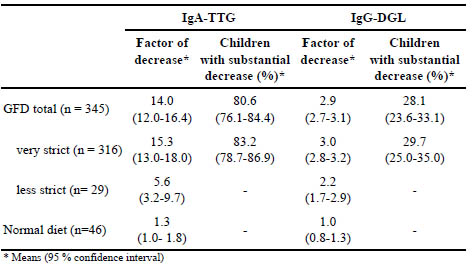
The major strengths of our study are the large number of patients prospectively
recruited, the central, blinded measurements of antibodies, and the serial dilution of
the blood samples so that even very high antibody concentrations could be measured
appropriately.
Conclusions
Measurement of IgA-TTG after 3 months can be useful in confirming the response to a
GFD, if initial concentrations are known. Blood samples have to be diluted enough to
determine the absolute antibody concentration. For comparability, it is necessary to
use of the same test kit for initial and follow-up assays. A decrease by more than a
factor of four can be considered a clear indication of a response to GFD. A smaller
decrease suggests a rather poor response, perhaps indicating low adherence to the diet
and need for clinical or dietetic advice. However, at least on the short-term, high IgATTG
concentrations at follow-up are consistent with good GFD response.
The concentration of IgG-DGL also decreases in the vast majority of cases, but does
not distinguish GFD response from random fluctuations as effectively as IgA-TTG. In
standard laboratory procedures, IgG-DGL does not reach the upper bound of the
measurement range very often, meaning that it could be a pragmatic albeit inferior
alternative to IgA-TTG.
References
1. Husby S, Koletzko S, Korponay-Szabó I, et al. European Society for Pediatric
Gastroenterology, Hepatology, and Nutrition guidelines for the diagnosis of coeliac
Disease. J Pediatr Gastroenterol Nutr 2012; 54: 136–160.
2. Wolf J, Hasenclever D, Petroff D, et al. Antibodies in the diagnosis of coeliac
disease: a biopsy-controlled, international, multicentre study of 376 children with
coeliac disease and 695 controls. PloS One 2014;9: e97853.
3. Wolf J, Petroff D, Richter T, et al. Validation of Antibody-based Strategies for
Diagnosis of Pediatric Celiac Disease Without Biopsy. Gastroenterology
2017;153: 410-419.e17.
4. Werkstetter KJ, Korponay-Szabó IR, Popp A, et al. Accuracy in Diagnosis of
Celiac Disease Without Biopsies in Clinical Practice. Gastroenterology 2017; 153:
924-935.
6 The role of intestinal microbiota in coeliac disease
Role of bacterial metabolism in gluten peptide
modification
Elena Verdú
Division of Gastroenterology, Department of Medicine, McMaster University,
Hamilton, Canada
No manuscript provided.
Gut microbiome trajectory and coeliac disease risk
Yolanda Sanz
Institute of Agrochemistry and Food Technology (IATA-CSIC), Valencia, Spain
No manuscript provided.
Influence of diets on microbiota from patients with noncoeliac
gluten sensitivity
Walburga Dieterich1,2, Detlef Schuppan3,4, Monic Schink1,2, Raphaela Schwappacher 1,2,
Stefan Wirtz1, Markus F. Neurath1, Yurdagül Zopf1,2
1 Medical Department 1, Friedrich-Alexander-Universität Erlangen-Nürnberg,
Germany,
2 Hector Center of Excellence for Nutrition, Exercise and Sports, University of
Erlangen-Nürnberg, Germany,
3 Institute of Translational Immunology, University Medical Center, Johannes-
Gutenberg University, Mainz, Germany,
4 Division of Gastroenterology, Beth Israel Deaconess Medical Center, Harvard
Medical School, Boston, MA, USA
Introduction
Cereals and especially wheat products are the main carbohydrate sources in Western
diets. Interestingly, in recent years patients more often complain about gastrointestinal
and extra-intestinal symptoms after eating gluten-containing cereal products. The main
symptoms are diarrhoea or obstipation, abdominal pain, flatulence, nausea, and
vomiting, as well as muscle, head and body aches, or fatigue. In this context, coeliac
disease is well characterised as a destructive intestinal immune response to
immunogenic cereal gluten peptides in genetically predisposed individuals, affecting
about 1 % of most populations worldwide [1]. In addition, non-coeliac gluten
sensitivity (NCGS) is now considered as a distinct clinical entity, caused by wheat
consumption [2,3]. In contrast to patients with wheat allergy or coeliac disease no
disease specific serological markers are known in patients with NCGS. Furthermore,
there is no mucosal damage in patients with NCGS, except of moderately increased
numbers of lymphocytes in some patients [3,4]. Since there are no reliable diagnostic
markers in NCGS, the prevalence data are vague and range from 0.6 % – 6 % [5,6].
Interestingly, recent double-blind placebo controlled studies showed that gluten trigger
the clinical symptoms only in a minority of patients and thus question the causative
role of gluten in NCGS [4,7].
In recent years, other elimination diets enjoyed great popularity. In this context, a
FODMAP-reduced diet (fermentable oligo-, di- and monosaccharides and polyols)
showed a clear improvement of clinical symptoms, especially in patients with IBS
[8,9], and a reduction in dietary FODMAPs also improved the gastrointestinal
symptoms in most patients with NCGS [7,10]. However, other wheat components like
amylase-trypsin inhibitors (ATI), which were shown to stimulate the native immune
system [11], or a microbial dysbiosis may also influence the pathogenesis of NCGS.
The aim of this study was to evaluate the effect of a low FODMAP diet and a gluten free diet (GFD) on the microbial composition from patients with NCGS compared
with microbiota pattern from healthy controls.
Materials and methods
Subjects
Nineteen patients with typical clinical presentation of NCGS were enrolled in the
study. All patients complained about intestinal and extra-intestinal symptoms which
appeared usually within one and twelve hours after ingestion of wheat products, and
symptoms resolved under gluten-free diet. Ten healthy controls without any diseases
and complaints on standard diets were enclosed. Informed consent was obtained from
all participants and the study protocol was approved by ethics committee of University
of Erlangen.
Patient characteristics and nutritional analysis
None of our participants adhered to a strict GFD at first presentation. Before study
beginning, the age, sex, medication, complaints and underlying diagnoses were
requested, and sera were analysed for routine blood parameters, vitamin levels and
micronutrients. A nutritional analysis using the nutrition software PRODI® (Nutri-
Science GmbH, Germany) was performed for analysis of proteins, fats and
carbohydrates.
Exclusion of wheat allergy and coeliac disease
Total and wheat-specific immunoglobulin E (IgE) were determined shortly before
study beginning (ImmunoCAP™250, ThermoFisher Scientific, Germany), and a prick
test was used to exclude cutaneous wheat sensitisation. Furthermore, coeliac specific
serum antibodies for deamidated gliadin and tissue transglutaminase were determined
from all study participants using commercially available kits (Eurospital S.p.A., Italy).
Genotyping from blood cells for HLA-DQ2 or -DQ8, were performed from all
participants according to manufacturer’s instructions (Eu-Gen, Eurospital S.p.A.,
Italy). Patients with NCGS underwent initial gastroduodenoscopy, and 4-5 duodenal
biopsies were taken to exclude coeliac disease.
Microbiome analysis
Stool samples were collected and stored at -20°C. Genomic bacterial DNA was
isolated from frozen samples with the QIAamp Fast DNA Stool Mini Kit (Qiagen,
Germany), and the V3-V4 region of bacterial 16sRNA genes was amplified, and DNA
was analysed on the Illumina MiSeq.
Study design
All participants ate a Western standard diet for at least 4 weeks before study onset. At
study start, the participants were instructed by a professional dietician to consume a low FODMAP diet for two weeks. After a transition period of at least five days, when
participants were requested to eat standard diet, the participants were instructed to
consume a GFD for another two weeks. Stool samples were collected under the
Western standard diet, and at the end of the low FODMAP and gluten-free diet,
respectively. Furthermore, patients completed questionnaires concerning nutrition,
gastrointestinal symptom rating score (GSRS), before study start and at the end of each
diet.
Statistics
Statistics were performed using GraphPad Prism 6 (GraphPad software Inc., USA).
Parametric t-test and non-parametric Mann-Whitney U were used to determine
statistical power. p<0.05 was considered statistically significant.
Results and discussion
Since the data from this study have not been published, only preliminary results are
presented here.
Laboratory and genetics
The laboratory and blood parameters for vitamins and micronutrients did not vary
between patients with NCGS and healthy controls.
The HLA-DQ2 or -DQ8 are mandatory to develop coeliac disease, but their role in
NCGS is still unclear. Interestingly, our data showed that these genotypes were overrepresented
in patients with NCGS (52.6 %), compared to controls (30 %) and
prevalence data in Western countries, and thus are in accordance with other reports
[3,4,12,13].
Dietary effects on clinical symptoms
Wheat containing standard diet caused clinical symptoms in all our patients with
NCGS. The main gastrointestinal complaints were flatulence, abdominal pain, and
diarrhoea. The severity of gastrointestinal symptoms was significantly different
between NCGS patients and healthy controls at the beginning of the study under
Western standard diet. Patients with NCGS reported a clear improvement of
gastrointestinal symptoms under a low FODMAP diet, and their complaints further
improved when they consumed the GFD.
Thus, our study demonstrated a substantial improvement of gastrointestinal symptoms
in patients with NCGS already after consuming a low FODMAP diet. Apart from the
risk that a non-blinded two-week FODMAP low diet may have caused some placebo
effect, our data confirmed the observations from others that FODMAPs at least
partially account for the clinical symptoms in NCGS patients [10,14]. However, gluten
elimination further improved and almost completely resolved clinical symptoms in all
our patients with NCGS, suggesting a multifactorial trigger in NCGS.
Preliminary data of dietary effects on microbiota
It was demonstrated that patients suffering from IBS profit from a low FODMAP diet
[8,15,16], but the mechanisms how a FODMAP-restricted diet may affect IBS
symptoms is still unclear. An important role of intestinal bacteria and bacterial
dysbiosis was suggested in IBS [17].
When we compared the microbiota on phylum level, there was a clear trend to
decreased Bacteroidetes and increased Firmicutes in NCGS patients compared to
healthy controls. Interestingly, these data resemble findings from other studies, where
a similar dysbalance was described in most IBS patients [18,19]. Whereas, dietary
changes had only a minor effect on microbiota in our controls, the diets caused
significant variations in our patients with NCGS.
Thus our study confirmed other reports that also showed a higher microbial fluctuation
in patients with IBS, and dietary changes resulted in more variations in microbial
composition, whereas healthy controls showed higher stability of their microbiota
[20,21]. Although there were major variations after dietary changes especially in
patients with NCGS, we noticed a substantial clustering of the individual microbiota,
even under the different diets. This is in accordance with a prior study showing that a
major dietary modification caused microbial shifts within 24 hours, while the
enterotypes with their dominant species remained stable during the whole ten-day
observation period [22].
Conclusions
Our data suggest a multifactorial ongoing in NCGS caused by FODMAPs, combined
with a gluten-related immune reaction and a microbial dysbalance. However, the direct
causal role of over- or under-represented microorganisms in NCGS remains unclear
and more detailed analyses are required.
References
1. Tack GJ, Verbeek WH, Schreurs MW, et al. The spectrum of celiac disease:
epidemiology, clinical aspects and treatment. Nat Rev Gastroenterol Hepatol 2010;
7: 204-213.
2. Catassi C, Elli L, Bonaz B, et al. Diagnosis of Non-Celiac Gluten Sensitivity
(NCGS): The Salerno Experts' Criteria. Nutrients 2015; 7: 4966-4977.
3. Volta U, Tovoli F, Cicola R, et al. Serological tests in gluten sensitivity (nonceliac
gluten intolerance). J Clin Gastroenterol 2012; 46: 680-685.
4. Zanini B, Basche R, Ferraresi A, et al. Randomised clinical study: gluten challenge
induces symptom recurrence in only a minority of patients who meet clinical
criteria for non-coeliac gluten sensitivity. Aliment Pharmacol Ther 2015; 42: 968-
976.
5. DiGiacomo DV, Tennyson CA, Green PH, et al. Prevalence of gluten-free diet
adherence among individuals without celiac disease in the USA: results from the
Continuous National Health and Nutrition Examination Survey 2009-2010. Scand J
Gastroenterology 2013; 48: 921-925.
6. Sapone A, Bai JC, Ciacci C, et al. Spectrum of gluten-related disorders: consensus
on new nomenclature and classification. BMC Med 2012; 10:13.
7. Biesiekierski JR, Peters SL, Newnham ED, et al. No effects of gluten in patients
with self-reported non-celiac gluten sensitivity after dietary reduction of
fermentable, poorly absorbed, short-chain carbohydrates. Gastroenterology 2013;
145: 320-328 e1-3.
8. Halmos EP, Power VA, Shepherd SJ, et al. A diet low in FODMAPs reduces
symptoms of irritable bowel syndrome. Gastroenterology 2014; 146: 67-75 e5.
9. Nanayakkara WS, Skidmore PM, O'Brien L, et al. Efficacy of the low FODMAP
diet for treating irritable bowel syndrome: the evidence to date. Clin Exp
Gastroenterology 2016; 9: 131-142.
10. van Gils T, Nijeboer P, CE IJ, et al. Prevalence and characterization of selfreported
gluten sensitivity in The Netherlands. Nutrients 2016; 8: 714.
11. Zevallos VF, Raker V, Tenzer S, et al. Nutritional Wheat Amylase-Trypsin
Inhibitors Promote Intestinal Inflammation via Activation of Myeloid Cells.
Gastroenterology 2017;152: 1100-13 e12.
12. Biesiekierski JR, Newnham ED, Irving PM, et al. Gluten causes gastrointestinal
symptoms in subjects without celiac disease: a double-blind randomized placebocontrolled
trial. Am J Gastroenterol 2011; 106: 508-514; quiz 15.
13. Carroccio A, D'Alcamo A, Cavataio F, et al. High proportions of people with
Nonceliac Wheat Sensitivity have autoimmune disease or antinuclear antibodies.
Gastroenterology 2015; 149: 596-603 e1.
14. Skodje GI, Sarna VK, Minelle IH, et al. Fructan, rather than gluten, induces
symptoms in patients with self-reported non-celiac gluten sensitivity.
Gastroenterology 2017; 154: 529-539.e2.
15. Bohn L, Storsrud S, Liljebo T, et al. Diet low in FODMAPs reduces symptoms of
irritable bowel syndrome as well as traditional dietary advice: a randomized
controlled trial. Gastroenterology 2015; 149: 1399-407 e2.
16. McIntosh K, Reed DE, Schneider T, et al. FODMAPs alter symptoms and the
metabolome of patients with IBS: a randomised controlled trial. Gut 2017; 66:
1241-51.
17. Galipeau HJ, Verdu EF. Gut microbes and adverse food reactions: Focus on gluten
related disorders. Gut Microbes 2014; 5: 594-605.
18. Jeffery IB, O'Toole PW, Ohman L, et al. An irritable bowel syndrome subtype
defined by species-specific alterations in faecal microbiota. Gut 2012; 61: 997-
1006.
19. Rajilic-Stojanovic M, Biagi E, Heilig HG, et al. Global and deep molecular
analysis of microbiota signatures in fecal samples from patients with irritable
bowel syndrome. Gastroenterology 2011; 141: 1792-801.
20. Halfvarson J, Brislawn CJ, Lamendella R, et al. Dynamics of the human gut
microbiome in inflammatory bowel disease. Nat Microbiol 2017; 2: 17004.
21. Rajilic-Stojanovic M, Jonkers DM, Salonen A, et al. Intestinal microbiota and diet
in IBS: causes, consequences, or epiphenomena? Am J Gastroenterol 2015; 110:
278-287.
22. Wu GD, Chen J, Hoffmann C, et al. Linking long-term dietary patterns with gut
microbial enterotypes. Science 2011; 334: 105-108.
7 Statements on current developments
concerning gluten analysis, clinical and legal
aspects
Comparison of analytical data from collaborative
studies using R5 and G12 antibodies
Paul Wehling
General Mills, Inc., Golden Valley, USA
No manuscript provided.
News from Codex and Regulatory Affairs
Hertha Deutsch
AOECS Codex Delegate, Austrian Coeliac Society, Vienna
Introduction
AOECS, the Association Of European Coeliac Societies, has Observer status in the
Codex Alimentarius Commission since 1992. Information about the organisation, the
duties, the Codex Standards and Guidelines, the Procedural Manual and all the
extensive work of all Codex Committees is published on their website [1].
Regarding gluten-free foods, especial two Codex Committees are very important:
Codex Committee on Nutrition and Foods for Special Dietary Uses (CCNFSDU) and
the Codex Committee on Methods of Analysis and Sampling (CCMAS).
Since November 2014, CCNFSDU and CCMAS have been considering the G12
method as a potential additional method for gluten determination.
Considerations in the past years
Details about the discussions in the CCNFSDU and CCMAS sessions were published
in the Proceedings of the Working Group on Prolamin Analysis and Toxicity [2,3].
Here are the results:
Conclusion of the CCNFSDU session in November 2014
“CCNFSDU36 agreed to ask CCMAS to examine ELISA G12 as a potential
additional method for determination of gluten.”
Reply from the CCMAS session in February 2015
“When considering “ELISA G12” as a potential additional method for inclusion in
CODEX STAN 118-1979, CCMAS 36 noted that any potential endorsement of G12
would be as a Type I procedure and that it would not be possible to have two Type I
methods in the Standard for the same matrices and determination. CCMAS further
observed that if the G12 method for detection of the toxic fraction in gluten harmful
for individuals were added, the provision in the Standard would need to be
differentiated to allow for both methods (R5 and G12) to be included as Type I
methods. The Committee noted that G12 had been validated for gluten-free foods, rice
matrices, whereas R5 had been validated for gluten-free foods, maize matrices.
CCMAS 36 recommended that decision in this regard should be taken by CCNFSDU.
The Committee is invited to consider the above reply.”
Conclusion of the CCNFSDU session in November 2015
“The Committee noted the reply from CCMAS in particular with respect to validation
of the R5 and G12 methods, based on the two matrices, maize and rice but questioned:
which method to adopt for mixed matrices; the comparability of the two methods (if
different results emerge) and the implications for “gluten-free” labelling. The
Committee decided to seek further clarification from CCMAS with the following
request:
Taking into account that the thresholds in CODEX STAN 118-1979 were
established on the basis of the results given by the ELISA R5 Method, can CCMAS
confirm that the results of the two methods (R5 and G12) are fully comparable for
all products covered by the standard, in particular:
̵ products manufactured from ingredients naturally free of gluten (e.g. buckwheat,
millet, amaranth, quinoa etc.);
̵ products manufactured from gluten-containing ingredients (e.g. partially
hydrolysed wheat protein, wheat starch, malt extract, glucose syrups etc.);
̵ products based on oats;
̵ liquid matrices.”
Reply from the CCMAS session in February 2016
“The Committee agreed to inform CCNFSDU that the two methods (R5 and G12) for
the determination of gluten are not comparable; that comparability data for the two
methods were not available; and mixed matrices are not included in the scope of either
of the methods obtained during their validation. The developers of these proprietary
methods might be able to provide further information on the applicability of the
methods.”
Recent CCNFSDU and CCMAS sessions
The 38th session of the Codex Committee on Nutrition and Foods for Special Dietary
Uses (CCNFSDU) took place from 5 - 9 December 2016 in Hamburg, Germany. The
Committee was attended by delegates from 56 Member Countries, one Member
Organisation and 38 International Organisations, one of them was AOECS.
At agenda item 2 „Matters referred to the Committee by the Codex Alimentarius
Commission and/or other subsidiary bodies“, sub-item „Matters for action“, the
CCNFSDU has to consider the reply from the CCMAS session in February 2016.
During the discussion I informed CCNFSDU that the Working Group on Prolamin
Analysis and Toxicity is working on comparability studies and the conclusion of this
item is in the CCNFSDU report [4]:
“The Committee agreed not to include the ELISA G12 method in the Standard for
Foods for Special Dietary Use for Persons Intolerant to Gluten (CODEX STAN 118-
1979) noting that there were no comparability results with R5; and it would be
considered at a future date when the results from the ongoing comparability studies by
the international Working Group on Prolamin Analysis and Toxicity become
available.”
The 38th session of the Codex Committee on Methods of Analysis and Sampling
(CCMAS) took place on 8 - 12 May 2017 in Budapest, Hungary. The Committee was
attended by 47 Member Countries, one Member Organisation and 11 Observer
Organisations, one of them was AOECS.
At agenda item 2 “Matters referred to the Committee by the Codex Alimentarius
Commission and/or other subsidiary bodies”, sub-item “Matters for information”, the
secretary informed CCMAS about the decision of CCNFSDU - see above. The
Committee noted this information without any comments.
Since a few years an important item is on the agenda of CCMAS:
Development of Procedures/Guidelines for determining equivalency to Type I
Methods
In the past years, an electronic Working Group considered in general the subject of
equivalency of Type 1 Methods and finally CCMAS agreed to the following
conclusion for the CCMAS report [5]:
“The Committee could not reach consensus on the use and scope of the equivalency
approach and agreed to reconsider this matter in the future when more information
became available. The Committee noted that most of the work in determining
equivalence falls on the Standards Development Organisations (SDOs), and noted the
offer of the SDOs, through the Inter-Agency Meeting (IAM), to look into this matter
and provide recommendations to a future session of CCMAS.”
Codex Methods of Analysis
The Codex Procedural Manual [1] defines 4 types of methods. If any new method
should become a Codex Method, it is mandatory to fit in one of the following types:
“(a) Defining Methods (Type I)
Definition: A method which determines a value that can only be arrived at in terms of
the method per se and serves by definition as the only method for establishing the
accepted value of the item measured.
(b) Reference Methods (Type II)
Definition: A Type II method is the one designated Reference Method where Type I
methods do not apply. It should be selected from Type III methods (as defined below).
It should be recommended for use in cases of dispute and for calibration purposes.
(c) Alternative Approved Methods (Type III)
Definition: A Type III Method is one which meets the criteria required by the
Committee on Methods of Analysis and Sampling for methods that may be used for
control, inspection or regulatory purposes.
(d) Tentative Method (Type IV)
Definition: A Type IV Method is a method which has been used traditionally or else
has been recently introduced but for which the criteria required for acceptance by the
Committee on Methods of Analysis and Sampling have not yet been determined.”
Conclusion
Gluten-free foods are imported and exported all over the globe. It is essential for
coeliacs, food manufacturers and national food control authorities that all approved
gluten detection methods should show comparable results.
References
1. Codex Alimentarius Commission http://www.codexalimentarius.org/
2. Deutsch H. News from Codex and regulatory affairs. In: Koehler P (ed):
Proceedings of the 29th Meeting of the Working Group on Prolamin Analysis and
Toxicity. Deutsche Forschungsanstalt für Lebensmittelchemie, Freising, 2016; pp.
117-122
3. Deutsch H. News from Codex and regulatory affairs. In: Koehler P (ed):
Proceedings of the 30th Meeting of the Working Group on Prolamin Analysis and
Toxicity. Deutsche Forschungsanstalt für Lebensmittelchemie, Freising, 2017; pp.
131-135
4. Report of the 38th session of the Codex Committee on Nutrition and Foods for
Special Dietary Uses, Hamburg, Germany, 5 - 9 December 2016
5. Report of the 38th session of the Codex Committee on Methods of Analysis and
Sampling, Budapest, Hungary 8 - 12 May 2017
8 Perspectives and action plan of the PWG
Peter Koehler
Biotask AG, Esslingen, Germany
The Prolamin Working Group executive meeting and joint discussion held on 29
September 2017, led to the decisions and statements outlined below.
Action plan
I. Analytical
Due to a job change of the chairman, the distribution of the PWG gliadin
reference material is no longer possible by Peter Koehler.
Efforts to find alternative distributors (R-Biopharm AG, Leibniz-Institute for
Food Systems Biology) failed.
The distribution of PWG-gliadin has been discontinued from October 2017
until a new distributor has been identified.
New reference material: The MoniQA initiative is currently identifying
cultivars of wheat, rye and barley and aims at providing flours as gluten
reference.
However, the PWG thinks that flour is not a suitable reference material and
supports a protein sample as reference material.
II. Clinical
For the symposium of the 2018 meeting the topic “Wheat Genomics” has been
selected. In addition, a speaker on “Gluten Epitopes and Interaction with
Antibodies” will be invited
III. Members, Policy
Katharina Scherf is a new member of the group.
Rudolf Valenta and Martin Stern left the group.
New group members will be identified by the group and two potential members
will be invited to the next meeting.
Fernando Chirdo is responsible for the website.
This printed, citable book (print run: 250 copies with ISBN number) was made
possible by funding of Dr. SCHÄR GmbH/Srl, (Burgstall, BZ, Italy). It will be
distributed among leaders of opinion in gluten analysis and clinical medicine.
An electronic version can be downloaded free of charge from the PWG website.
Next meeting: 2018
We are very pleased to announce the venue for our meeting in 2018:
Ayr, Scotland, United Kingdom
Host:
Ms. Pauline Titchener
Neogen Europe Ltd.
E-mail: p.titchener@neogeneurope.com
Time: 27 - 29 September 2018
Focus of the meeting:
Wheat genomics
Gluten epitopes and interaction with antibodies
The meeting will be limited to 55 participants and attendance is by
invitation only. Invitations will be sent by March 2018. Registration
deadline will be June 15, 2018.
For registration please contact:
Ms. Pauline Titchener
Neogen Europe Ltd.
The Dairy School, Auchincruive
AYR, SCOTLAND, UK, KA6 5HU
Phone: +44 1292 525 600
Fax: +44 1292 525 601
E-mail: p.titchener@neogeneurope.com
Very special thanks to the host of this kind invitation!
|
|
List of Participants
GROUP MEMBERS
Prof. Dr. Fernando G. Chirdo
Universidad Nacional de La Plata
Facultad de Ciencias Exactas
Instituto de Estudios Immunologicos y
Fisiopatologicos - IIFP
Calle 47 y 115
(1900) LA PLATA, ARGENTINA
Phone: +54 221 423 5 333 (Int 45)
Fax: +54 221 422 6947
E-mail: fchirdo@biol.unlp.edu.ar
Prof. Dr. Paul J. Ciclitira
GSTT NHS Trust
Curve Business Hub, Unit 1
30B Wilds Rents
SE1 4QG LONDON
UNITED KINGDOM
Phone: +44 203 751 1104
Fax: +44 207 4033437
E-mail: pjcgastro@gmail.com
Prof. Dr. Carlo Catassi
(not attending), substituted by
Dr. Anil K. Verma
Università Politecnica delle Marche
Celiac Disease Research Laboratory
Dept of Pediatrics
Via Cadorna 5
60123 ANCONA, ITALY
Phone: +39 33 4292 2991
E-mail: anilkrvermaa@gmail.com
Prof. Dr. Conleth Feighery, MD
University of Dublin, Department of
Immunology, St. James’s Hospital
James’s Street
DUBLIN 8, IRELAND
Phone: +353 879969041
E-mail: con.feighery@tcd.ie
Prof. Dr. Peter Koehler
Biotask AG
Schelztorstraße 54-56
73728 ESSLINGEN, GERMANY
Phone: +49 711 31059068
Fax: +49 711 31059070
E-mail: peter.koehler@biotask.de
Prof. Dr. Knut Lundin
University of Oslo
Institute of Clinical Medicine
Postboks 1171, Blindern
0881 OSLO, NORWAY
Phone: +47 90980325
Fax: +47 23072410
E-mail: knut.lundin@medisin.uio.no
Prof. Dr. Thomas Mothes
Institut für Labormedizin der Universität
Leipzig
Liebigstraße 27
04103 LEIPZIG, GERMANY
Phone: +49 1577 3327893
Fax: +49 341 97 22373
E-mail: mothes@medizin.uni-leipzig.de
Dr. Katharina Scherf
Leibniz-Institut für Lebensmittel-
Systembiologie an der Technischen
Universität München
Lise Meitner-Straße 34
85354 FREISING, GERMANY
Phone: +49 8161712927
Fax: +49 8161712970
E-mail: k.scherf.leibniz-lsb@tum.de
Prof. Dr. Dr. Detlef Schuppan
I. Medizinische Klinik und Poliklinik
Universitätsmedizin der Johannes
Gutenberg-Universität Mainz
Institut für Translationale Medizin
Langenbeckstraße 1
55131 MAINZ, GERMANY
Phone: +49 6131 177355/177356/177104
Fax: +49 6131 177357
E-mail: detlef.schuppan@unimedizinmainz.de
Dr. René Smulders
Wageningen University & Research,
Plant Research
Droevendaalsesteeg 1
6708 PB WAGENINGEN,
THE NETHETRLANDS
Phone: +31 620298266
E-mail: rene.smulders@wur.nl
Dr. Olivier Tranquet
INRA
Rue de la Géraudière BP 71627
44316 NANTES CEDEX 3, FRANCE
Phone: +33 2 40675027
Fax: +33 240675025
E-mail: olivier.tranquet@inra.fr
Prof. Dr. Riccardo Troncone
Department of Pediatrics and European
Laboratory for the Investigation of
Food-induced Diseases
Department of Padiatrics
Via Pansini 5
80131 NAPLES, ITALY
Phone: +39 3483132274
Fax: +39 0817463116
E-mail: troncone@unina.it
HOSTS
Dr. Markus Brandt
Böcker Sauerteig GmbH & Co. KG
Ringstraße 55-57
32427 MINDEN, GERMANY
Phone: +49 571 8379943
Fax : +49 571 8379920
E-mail: markus.brandt@sauerteig.de
Mr. Markus Düsterberg
Böcker Sauerteig GmbH & Co. KG
Ringstraße 55-57
32427 MINDEN, GERMANY
Phone: +49 571 837990
E-mail: markus.duesterberg@sauerteig.de
Ms. Karoline Schreiber
Böcker Sauerteig GmbH & Co. KG
Ringstraße 55-57
32427 MINDEN, GERMANY
Phone: +49 571 837990
E-mail: karoline.schreiber@sauerteig.de
INVITED SPEAKERS
Prof. Dr. Yolanda Sanz
Institute of Agrochemistry and Food
Technology (IATA-CSIC)
Avenida Agustin Escardino 7
46980 PATERNA, VALENCIA,
SPAIN
Phone: +34 963 900022 2128
E-mail: yolsanz@iata.csic.es
Dr. Walburga Dieterich
Universitätsklinik Erlangen,
Medizinische Klinik 1
Ulmenweg 18
91054 ERLANGEN, GERMANY
Phone: +49 9131 8535128
E-Mail:
walburga.dieterich@uk-erlangen.de
Prof. Dr. Elena Verdú
McMaster University
Health Sciences Centre
1280 Main Street West
L8S 4K1, HAMILTON, CANADA
Phone: +1 905 525 9140
E-mail: verdue@mcmaster.ca
GUESTS
Dr. Twan America
Wageningen University & Research,
Plant Research
Droevendaalsesteeg 1
6708 PB WAGENINGEN,
THE NETHERLANDS
Phone: +31 317 483050
E-mail: twan.america@wur.nl
Mrs. Güler Atasoy
Mersin University
Soli sitesi G blok kat:3 No:308
33190 MERSIN, TURKEY
Phone: +90 537 2681058
E-mail: gleratasoy@gmail.com
Mrs. Sofia Beisel
Deutsche Zöliakiegesellschaft e.V.
Kupferstraße 36
70565 STUTTGART, GERMANY
Phone: +49 711 45998115
Fax: 49 711 459981 50
E-mail: sofia.beisel@dzg-online.de
Mrs. Zsofía Birinyi
MTA Centre of Agricultural Research
Department of Applied Genomics
Brunszvik u 2
2462 MARTONVÁSÁR, HUNGARY
Phone: +36 225 69521
Fax: +36 225 69514
E-mail: birinyi.zsofia@agrar.mta.hu
Dr. Georg Böcker
Ernst Böcker GmbH & Co KG
Ringstraße 55-57
32427 MINDEN, GERMANY
Phone: +49 571 837990
Fax: +49 571 8379920
E-mail: gb@sauerteig.de
Mrs. Helen Brown
Campden BRI Ltd
Station Road
GL55 6LD CHIPPING CAMPDEN,
UNITED KINGDOM
Phone: +44 1386 842016
Fax: +44 1386 842100
E-mail: helen.brown@campdenbri.co.uk
Dr. Angel Cebolla-Ramirez
Biomedal, SL
Av. Américo Vespucino n° 5
- Bloque 4 - 1a planta -
41092 SEVILLA, SPAIN
Phone: +34 954 081276
Fax: +34 954 081279
E-mail: acebolla@biomedal.com
Dr. Michelle Colgrave
CSIRO Agriculture and Food
306 Carmody Rd, St Lucia
4067 BRISBANE, AUSTRALIA
Phone: +61 42 2670005
Fax: +61 732142900
E-mail: michelle.colgrave@csiro.au
Dr. Johan De Meester
Cargill R&D Centre Europe
Havenstraat 84
B-1800 VILVOORDE, BELGIUM
Phone: +32 225 70733
E-mail: Johan_De_Meester@cargill.com
Mrs. Hertha Deutsch
Österreichische Arbeitsgemeinschaft
Zöliakie
Anton Baumgartner Straße 44/C5/2302
1230 VIENNA, AUSTRIA
Phone: +43 166 71887
E-mail: hertha.deutsch@chello.at
Mr. Richard Fielder
Bio-Check (UK)
Spectrum House, Llys Edmund Prys,
St. Asaph Business Park
LL17 0JA ST. ASAPH,
UNITED KINGDOM
Phone: +44 1745 335165
Fax: +44 1745 582867
E-mail: richard@biocheck.uk.com
Dr. Gyöngyvér Gell
MTA Centre of Agricultural Research
Department of Applied Genomics
Brunszvik u 2
2462 MARTONVÁSÁR, HUNGARY
Phone: +36 225 69521
Fax: +36 225 69514
E-mail: gell.gyongyver@agrar.mta.hu
Mrs. Anna Gibert Casamada
Associació Celíacs de Catalunya
Independencia, 257
08026 BARCELONA, SPAIN
Phone: +34 934 121789
E-mail: annagibertc@gmail.com
Dr. Thomas Grace
Bia Diagnostics/Elution Technologies
480 Hercules Dr.
5446 COLCHESTER, VT, USA
Phone: +1 802 540 0148
Fax: +1 8025400147
E-mail:
thomasgrace@biadiagnostics.com
Mrs. Robin Grace
Bia Diagnostics/Elution Technologies
480 Hercules Dr.
5446 COLCHESTER, VT, USA
Phone: +1 802 540 0148
E-mail: robin@biadiagnostics.com
Mrs. Mia Hallgren
Livsmedelsverket, National Food
Agency Sweden
Hamnesplanaden 5, Box 522
751 26 UPPSALA, SWEDEN
Phone:+46 18 175342
E-mail: Mia.Hallgren@slv.se
Mrs. Katri Hautanen
Fria Bröd AB
Fältspatsgatan 10
42130 FRÖLUNDA, SWEDEN
Phone: +46 705156683
E-mail: katri.hautanen@fria.se
Dr. Xin Huang
University of Helsinki
Department of Food and
Environmental Sciences
Agnes Sjöbergin katu 2, PL66
00014 HELSINKI, FINLAND
Phone: +358 451210203
E-mail: xin.huang@helsinki.fi
Dr. Götz Kröner
Hermann Kröner GmbH
Lengericher Straße 158
49479 IBBENBÜREN, GERMANY
Phone: +49 5451 9447 11
Fax: +49 5451 9447 811
E-mail: kroener@kroener-staerke.de
Ms. Lilian Kuster
Romer Labs Division Holding GmbH
Erber Campus 1
3131 GETZERSDORF, AUSTRIA
Phone: +43 2782 8030
E-mail: lilian.kuster@romerlabs.com
Ms. Stella Lindeke
R-Biopharm AG
An der neuen Bergstraße 17
64297 DARMSTADT, GERMANY
Phone: +49 6151 8102 92
E-mail: s.lindeke@r-biopharm.de
Mrs. Maria Jose Martinez Esteso
Institute for Reference Materials and
Measurements, Joint Research Centre,
European Commission
Retiesweg 111
2440 GEEL, BELGIUM
E-mail: maria-jose.martinezesteso@ec.europa.eu
Mrs. Ombretta Polenghi
Dr. Schär R&D Centre
c/o AREA Science Park
Padriciano, 99
I-34149 TRIESTE, ITALY
Phone: +39 040 3755 381
E-mail:
ombretta.polenghi@drschaer.com
Dr. Elena Quesada Hernández
Biomedal, SL
Av. Américo Vespucino n° 5
- Bloque 4 - 1a planta -
41092 SEVILLA, SPAIN
Phone: +34 954 08 12 76
Fax: +34 954 08 12 79
E-mail: elena.quesada@biomedal.com
Mrs. Catherine Remillieux-Rast
Association Française des Intolérants
au Gluten (AFDIAG)
23 Rue de Venise
78740 VAUX-SUR-SEINE, FRANCE
Phone: +33 681270911
Fax: +33 130993668
E-mail: c.remillieux_rast@yahoo.fr
Dr. Adrian Rogers
Romer Labs UK Ltd.
The Health Business and
Technical Park
WA74QX RUNCORN, CHESHIRE,
UNITED KINGDOM
Phone: +44 845519 0510
E-mail: adrian.rogers@romerlabs.com
Mr. Nermin Sajic
EuroProxima B.V.
Beijerinckweg 18
6827 BN ARNHEM,
THE NETHERLANDS
Phone: +31 263630364
E-mail: nermin.sajic@europroxima.com
Mrs. Tanja Schirmer
Leibniz-Institut für Lebensmittel-
Systembiologie an der Technischen
Universität München
Lise Meitner-Straße 34
85354 FREISING, GERMANY
Phone: +49 8161712941
Fax: +49 8161712970
E-mail: t.schirmer.leibniz-lsb@tum.de
Dr. Juan Ignacio Serrano-Vela
Asociación de Celíacos y Sensibles Al
Gluten, Comunidad de Madrid
Calle Lanuza 19-bajo
28028 MADRID, SPAIN
Phone: +34 917130147
Fax: +34 917258059
E-mail: nachoserrano@celiacosmadrid.org
Prof. Dr. Edurne Simón
University of the Basque Country
UPV/EHU
Paseo de la Universidad, 7
1006 VITORIA-GASTEIZ, SPAIN
Phone: +34 945013069
Fax: +34 945013014
E-mail: edurne.simon@ehu.eus
Ms. Pauline Titchener
Neogen Europe Ltd.
The Dairy School, Auchincruive
KA6 5HU AYR, SCOTLAND,
UNITED KINGDOM
Phone: +44 1292 525 600
Fax: +44 1292 525 601
E-mail: p.titchener@neogeneurope.com
Dr. Paul Wehling
General Mills, Inc.
9000 Plymouth Ave N
55427 GOLDEN VALLEY, USA
Phone: +1 763 764 4360
E-mail: paul.wehling@genmills.com
Dr. Thomas Weiss
R-Biopharm AG
An der neuen Bergstraße 17
64297 DARMSTADT, GERMANY
Phone: +49 6151 8102 186
E-mail: t.weiss@r-biopharm.de
Dr. Nathalie Widmann
R-Biopharm AG
An der neuen Bergstraße 17
64297 DARMSTADT, GERMANY
Phone: +49 6151 8102 7220
E-mail: n.widmann@r-biopharm.de
Mrs. Maren Wiese
Hermann Kröner GmbH
Lengericher Straße 158
49479 IBBENBÜREN, GERMANY
Phone: +49 5451 9447 12
E-mail: wiese@kroener-staerke.de
Ms. Rebecca Williams
Bio-Check (UK)
Spectrum House, Llys Edmund Prys,
St. Asaph Business Park
LL17 0JA ST. ASAPH,
UNITED KINGDOM
Phone: +44 1745 335 165
E-mail: rebecca@biocheck.uk.com
Ms. Stephanie Zühlke
K. G. Jebsen-senter for cøliakiforskning
Immunologisk institutt, Rikshospitalet
Sognsvannsveien 20
0372 OSLO, NORWAY
Phone: +47 46523358
E-mail: stephanie.zuhlke@medisin.uio.no
Impressum
Proceedings of the 31th Meeting
WORKING GROUP
on PROLAMIN ANALYSIS and TOXICITY
28 - 30 September 2017
Minden, Germany
This work including all parts is subject to copyright. All rights are reserved and any
utilisation is only permitted under the provisions of the German Copyright Law.
Permissions for use must always be obtained from the publisher. This is in particular
valid for reproduction, translation, conversion to microfilm and for storage or
processing in electronic systems.
Scientific Organisation
Prof. Dr. Peter Koehler
biotask AG
Schelztorstraße 54-56, 73728 ESSLINGEN, GERMANY
Phone: +49 711 31059068; Fax: +49 711 31059070
E-mail: peter.koehler@biotask.de
Host
Dr. Markus Brandt
Head Research & Development
Ernst Böcker GmbH & Co KG
Ringstraße 55-57, 32427 MINDEN, GERMANY
Phone: +49 571 8379943; Fax: +49 571 8379920
E-mail: markus.brandt@sauerteig.de
Cover picture* and picture of participants:
Thomas Mothes
© Peter Koehler 2018
ISBN: 978-3-00-058802-0
|


